NC, NETCDF
Type of resources
Available actions
Topics
Keywords
Contact for the resource
Provided by
Years
Formats
-
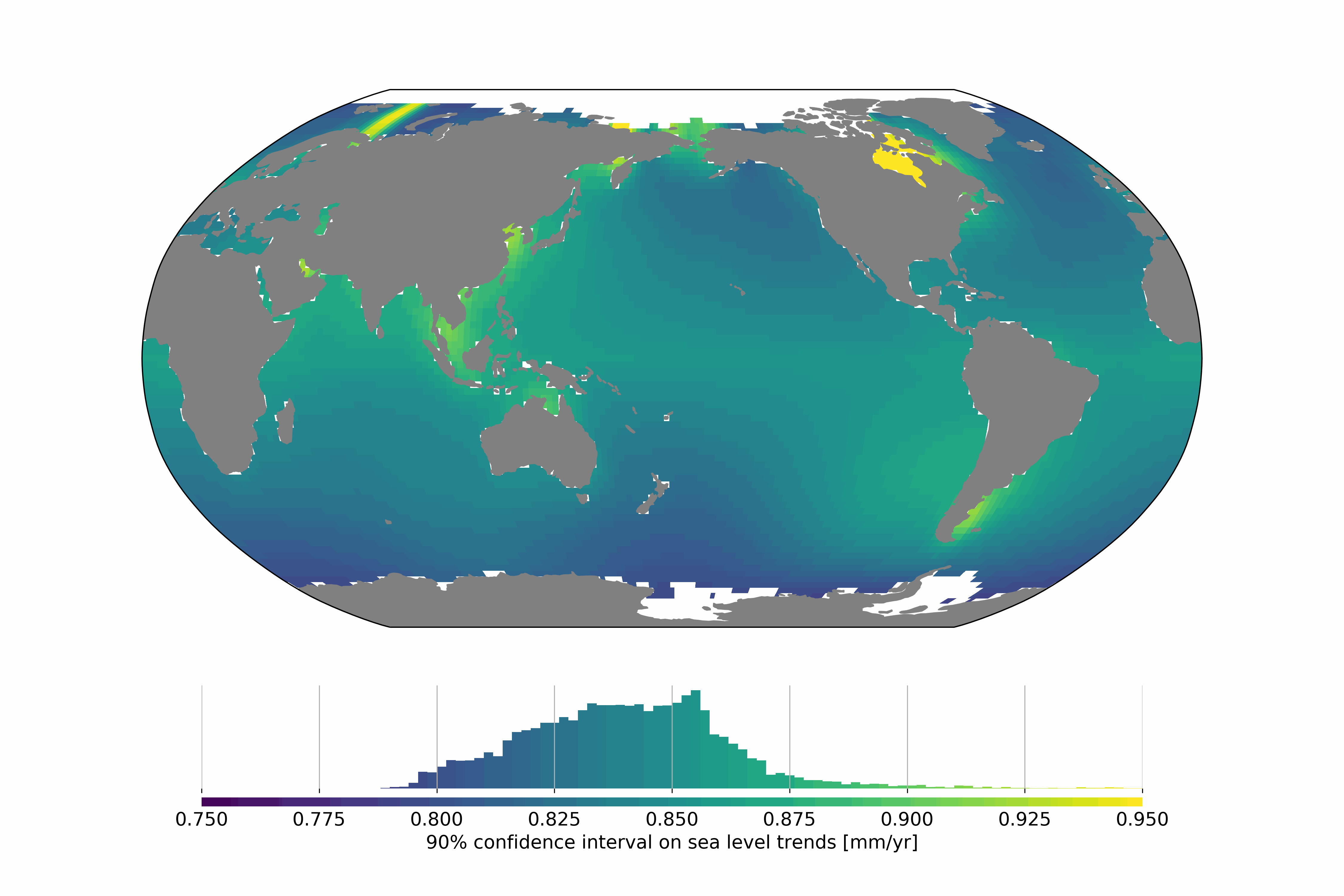
Satellite altimetry missions provide a quasi-global synoptic view of sea level over more than 25 years. The satellite altimetry constellation is used to build sea level maps and regional sea level indicators such as trends and accelerations. Estimating realistic uncertainties on these quantities is crucial to address some current climate science questions such as climate change detection and attribution or regional sea level budget closure for example. Previous studies have estimated the uncertainty for the global mean sea level (GMSL), but no uncertainty information is available at regional scales. In this study we estimate a regional satellite altimetry error budget and use it to derive maps of confidence intervals for local sea rise rates and accelerations. We analyze 27 years of satellite altimetry maps and derive the satellite altimetry error variance-covariance matrix at each grid point, prior to the estimation of confidence intervals on local trends and accelerations at the 90% confidence level using extended least squares estimators. Over 1993–2019, we find that the average local sea level trend uncertainty is 0.83 mm.yr-1 with local values ranging from 0.78 to 1.22 mm.yr-1. For accelerations, uncertainties range from 0.057 to 0.12 mm.yr-2, with a mean value of 0.063 mm.yr-2. Change history: - 2020/07/08: initial dataset submission over 1993-2018 - 2020/10/21: 1993-2019 update and addition of error levels
-
The glider operations in the MOOSE network started to be deployed regularly in 2010 in the North Western Mediterranean Sea, thanks to the setup of national glider facilities at DT-INSU/Ifremer (http://www.dt.insu.cnrs.fr/gliders/gliders.php) and with the support of the European project FP7-PERSEUS. Two endurance lines are operated: MooseT00 (Nice-Calvi; Ligurian Sea) and MooseT02 (Marseille-Menorca; Gulf of Lion). The all dataset here corresponds to raw data in the EGO format.
-

The continuously updated version of Copernicus Argo floats realtime currents product is distributed from Copernicus Marine catalogue: - https://resources.marine.copernicus.eu/?option=com_csw&view=details&product_id=INSITU_GLO_UV_NRT_OBSERVATIONS_013_048 The Argo current product generated by Copernicus in situ TAC is derived from the original trajectory data from Argo GDAC (Global Data Assembly Center) available at: - Argo float data and metadata from Global Data Assembly Centre (Argo GDAC). SEANOE. https://doi.org/10.17882/42182 In 2021, the GDAC distributes data from more than 15,000 Argo floats. Deep ocean current is calculated from floats drift at parking depth, surface current is calculated from float surface drift. An Argo float drifts freely in the global ocean, performing regular observation cycles. An observation cycle usually spreads over 10 days : - a surface descent to a parking depth (generally 1500 meters deep) - a 10-day drift at this parking depth - an ascent to the surface (vertical profile) - A short surface drift for data transmission The data transmitted at each cycle contain temperature, salinity observations (and additional biogeochemical parameters if applicable), positions (gps or argos), technical data. The ocean current product contains a NetCDF file for each Argo float. It is updated daily in real time by automated processes. For each cycle it contains the surface and deep current variables: - Date (time, time_qc) - Position (latitude, longitude, position_qc) - Pressure (pres, pres_qc, representative_park_pressure for parking drift, 0 decibar for surface drift) - Current (ewct, ewct_qc, nsct, nsct_qc; the current vector is positioned and dated at the last position of the N-1 cycle) - Duration (days) of the current variable sampling (time_interval) - Grounded indicator - Positions and dates have a QC 1 (good data). Positions and dates that do not have a QC 1 are ignored. The positions are measured during the surface drift (Argos or GPS positioning). For the deep current of cycle N, we take the last good position of cycle N-1 and the first good position of cycle N. For the surface current of cycle N, we take the first and last good position of the N cycle.
-
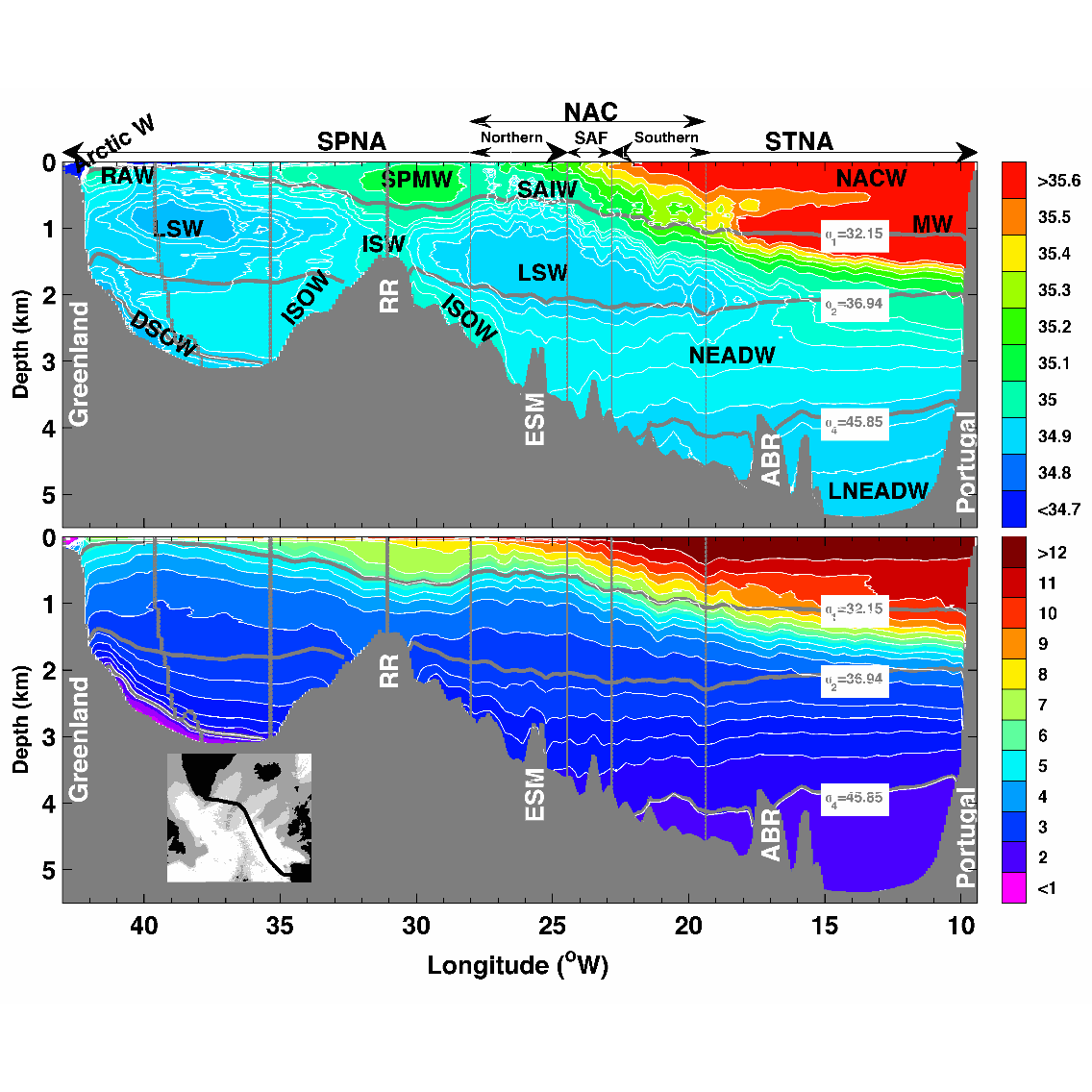
This data set contains the gridded hydrographic and transport data for the biennial Go-Ship A25 Greenland–Portugal OVIDE section from 2002 to 2012. The properties and transports are mapped on a 7km x 1m grid. Using a common grid facilitates the comparison between the different occupations of the line and the averaging. This data set was used in Daniault et al. (2016, Progress in Oceanography) to which the reader is referred for a description of the gridding method.
-
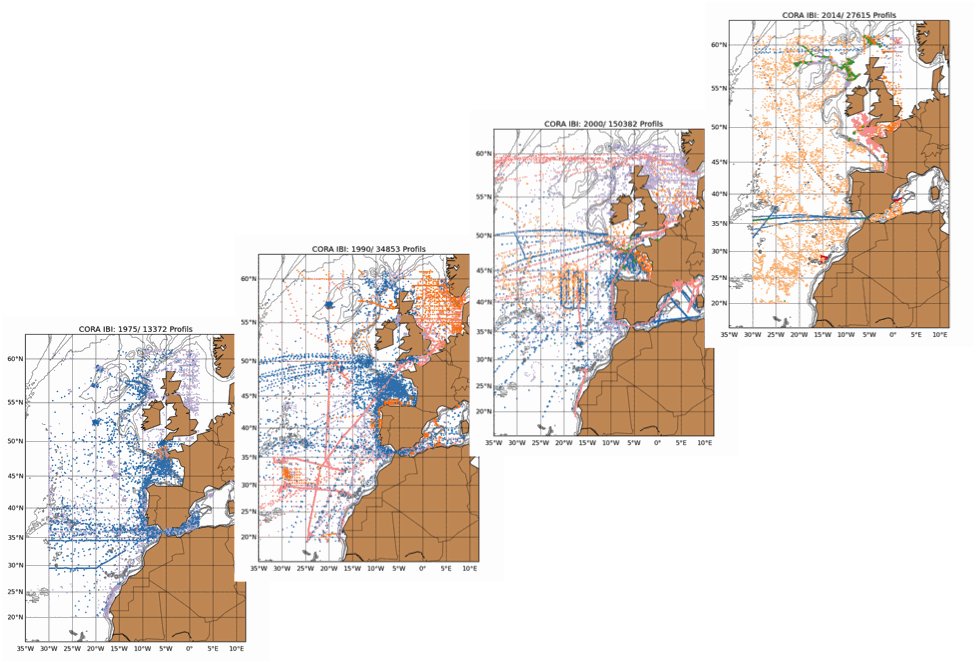
The Coriolis Ocean Dataset for Reanalysis for the Ireland-Biscay-Iberia region (hereafter CORA-IBI) product is a regional dataset of in situ temperature and salinity measurements. The latest version of the product covers the period 1950-2014. The CORA-IBI observations comes from many different sources collected by Coriolis data centre in collaboration with the In Situ Thematic Centre of the Copernicus Marine Service (CMEMS INSTAC). The observations integrated in the CORA-IBI product have been acquired both by autonomous platforms (Argo profilers, fixed moorings, gliders, drifters, sea mammals, fishery observing system from the RECOPESCA program), research or opportunity vessels ( CTDs, XBTs, ferrybox). This CORA-IBI product has been controlled using an objective analysis (statistical tests) method and a visual quality control (QC). This QC procedure has been developed with the main objective to improve the quality of the dataset to the level required by the climate application and the physical ocean re-analysis activities. It provides T and S individual profiles on their original level with QC flags. The reference level of measurements is immersion (in meters) or pressure (in decibars). It is a subset on the IBI (Iberia-Bay-of-Biscay Ireland) of the CMEMS product referenced hereafter. The main new features of this regional product compared with previous global CORA products are the incorporation of coastal profiles from fishery observing system (RECOPESCA programme) in the Bay of Biscay and the English Channel as well as the use of an historical dataset collected by the Service hydrographique de la Marine (SHOM).
-
C-RAID: Comprehensive Reprocessing of Drifting Buoy Data (1979-2018) The C-RAID (Copernicus - Reprocessing of Drifting Buoys) project delivers a comprehensive global reprocessing of historical drifting buoy data and metadata, providing climate-quality observations for marine and atmospheric research. Dataset Overview The C-RAID dataset encompasses metadata from 21 858 drifting buoys deployed between 1979 and 2018. Of these, 17 496 buoys have undergone complete reprocessing with scientific validation in delayed mode, including comparison against ERA5 reanalysis. Project Context Managed by the WMO DBCP Drifting Buoys Global Data Assembly Centre (GDAC) through Ifremer, Météo-France, and Ocean Sciences Division of Fisheries and Oceans Canada, C-RAID focuses on enhanced quality control and delivery of climate-quality drifting buoy data for the Marine Climate Data System (MCDS). Objectives - Complete reprocessing and clean-up of the historical drifting buoy data archive - Recovery and rescue of missing datasets - Reprocessing of Argos data with improved positioning using Kalman filter algorithms - Homogenization of quality control procedures across marine and atmospheric parameters Funding & Governance C-RAID was funded by the Copernicus Programme through the European Environment Agency (Contract # EEA/IDM/15/026/LOT1), supporting cross-cutting coordination activities for the in-situ component of Copernicus Services. Stakeholders & Partnerships The project is led by the DB-GDAC consortium (Ifremer, Météo-France) in collaboration with EUMETNET's E-SURFMAR programme, NOAA AOML, and JCOMMOPS. Key Achievements - Reprocessing of approximately 24 000 buoy-years of observations - Recovery of missing datasets and metadata through data rescue efforts - Implementation of homogeneous, rich metadata and data formats - Enhanced Argos location accuracy using Kalman filter reprocessing - Standardized quality control and validation procedures Data Access & FAIR Principles C-RAID provides FAIR (Findable, Accessible, Interoperable, Reusable) data access through: - Web-based data discovery portal for human users - API services for data discovery, subsetting, and download (machine-to-machine access) Target Users The dataset serves major operational and research programmes including: - Copernicus Climate Change Service (C3S) - Copernicus Marine Environment Monitoring Service (CMEMS) - iQuam (in-situ SST Quality Monitor) - ICOADS (International Comprehensive Ocean-Atmosphere Data Set) - GHRSST (Group for High Resolution Sea Surface Temperature) - ISPD (International Surface Pressure Databank) - ICDC (Integrated Climate Data Center)
-
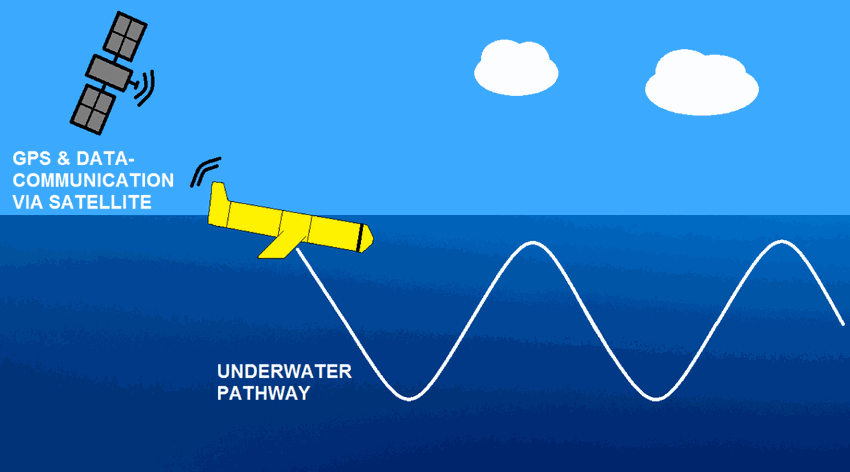
The observations of campe glider on imedia deployment (Mediterranean Sea - Western basin) are distributed in 4 files: - EGO NetCDF time-series (data, metadata, derived sea water current) - NetCDF profiles extracted from the above time-series - Raw data - JSON metadata used by the decoder The following parameters are provided : - Practical salinity - Sea temperature in-situ ITS-90 scale - Electrical conductivity - Sea water pressure, equals 0 at sea-level
-
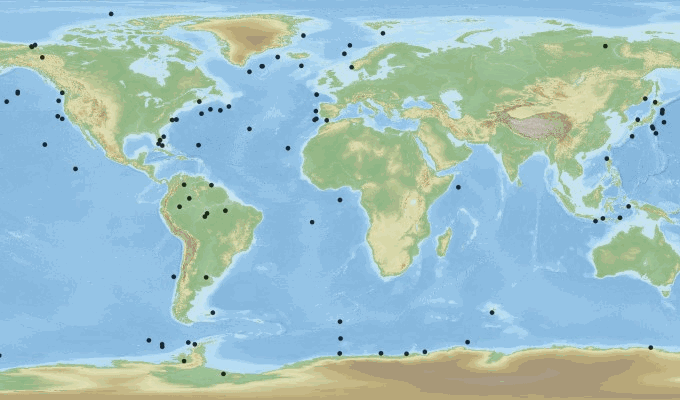
This product contains observations and gridded files from two up-to-date carbon and biogeochemistry community data products: Surface Ocean Carbon ATlas SOCATv2023 and GLobal Ocean Data Analysis Project GLODAPv2.2023. The SOCATv2023-OBS dataset contains >25 million observations of fugacity of CO2 of the surface global ocean from 1957 to early 2023. The quality control procedures are described in Bakker et al. (2016). These observations form the basis of the gridded products included in SOCATv2023-GRIDDED: monthly, yearly and decadal averages of fCO2 over a 1x1 degree grid over the global ocean, and a 0.25x0.25 degree, monthly average for the coastal ocean. GLODAPv2.2023-OBS contains >1 million observations from individual seawater samples of temperature, salinity, oxygen, nutrients, dissolved inorganic carbon, total alkalinity and pH from 1972 to 2021. These data were subjected to an extensive quality control and bias correction described in Olsen et al. (2020). GLODAPv2-GRIDDED contains global climatologies for temperature, salinity, oxygen, nitrate, phosphate, silicate, dissolved inorganic carbon, total alkalinity and pH over a 1x1 degree horizontal grid and 33 standard depths using the observations from the previous major iteration of GLODAP, GLODAPv2. SOCAT and GLODAP are based on community, largely volunteer efforts, and the data providers will appreciate that those who use the data cite the corresponding articles (see References below) in order to support future sustainability of the data products.
-
The DBCP – Data Buoy Cooperation Panel - is an international program coordinating the use of autonomous data buoys to observe atmospheric and oceanographic conditions, over ocean areas where few other measurements are taken. DBCP coordinates the global array of 1 600 active drifting buoys (August 2020) and historical observation from 14 000 drifting buoys. Data and metadata collected by drifting buoys are publically available in near real-time via the Global Data Assembly Centers (GDACs) in Coriolis-Ifremer (France) and MEDS (Canada) after an automated quality control (QC). In long term, scientifically quality controlled delayed mode data will be distributed on the GDACs. Disclaimer: the DB-GDAC is under construction. It is currently (January 2020) aggregating data from the Coriolis DAC (E-Surfmar, Canada). Additional DACs are considered. An interim provision from GTS real-time data to GDAC may be provided from Coriolis DAC.
-
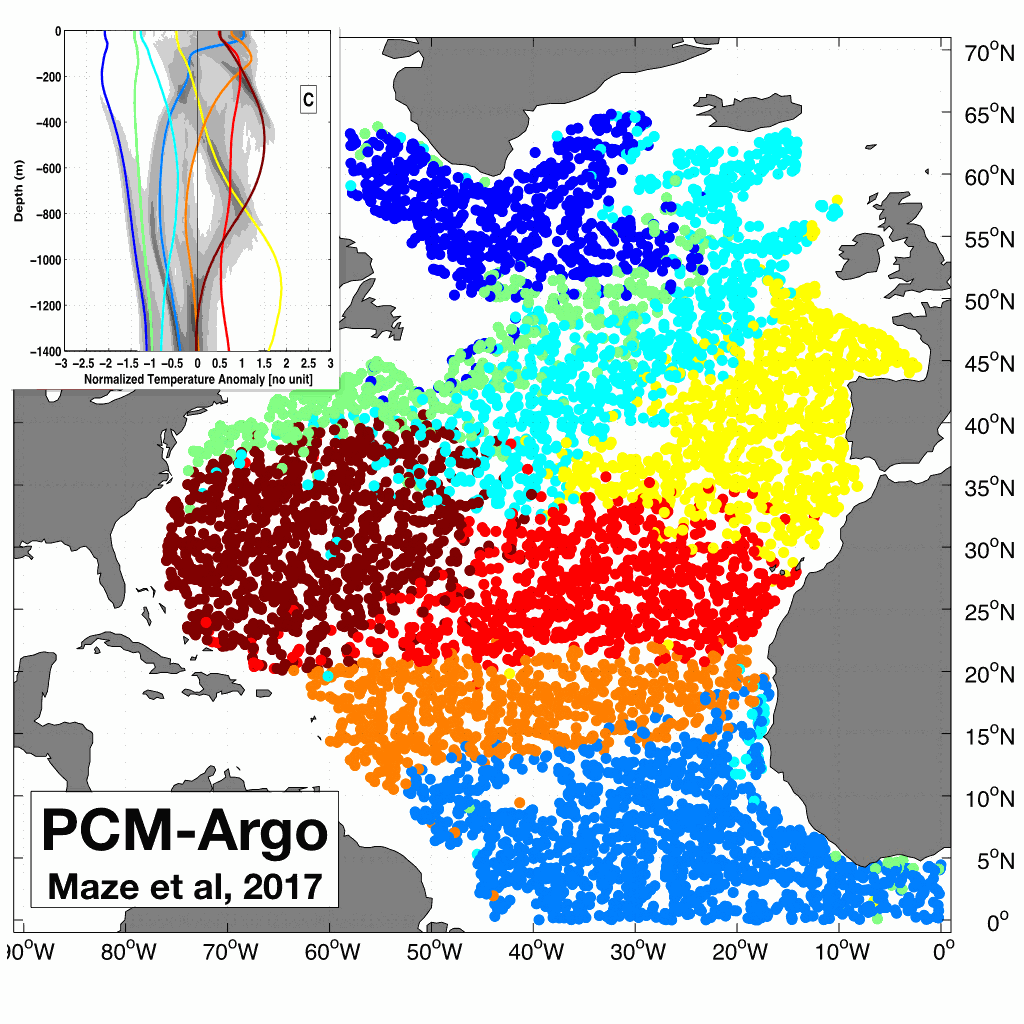
A quantitative understanding of the integrated ocean heat content depends on our ability to determine how heat is distributed in the ocean and what are the associated coherent patterns. This dataset contains the results of the Maze et al., 2017 (Prog. Oce.) study demonstrating how this can be achieved using unsupervised classification of Argo temperature profiles. The dataset contains: - A netcdf file with classification~results (labels and probabilities) and coordinates (lat/lon/time) of 100,684 Argo temperature profiles in North Atlantic. - A netcdf file with a Profile Classification Model (PCM) that can be used to classify new temperature profiles from observations or numerical models. The classification method used is a Gaussian Mixture Model that decomposes the Probability Density Function of the dataset into a weighted sum of Gaussian modes. North Atlantic Argo temperature profiles between 0 and 1400m depth were interpolated onto a regular 5m grid, then compressed using Principal Component Analysis and finally classified using a Gaussian Mixture Model. To use the netcdf PCM file to classify new data, you can checkout our PCM Matlab and Python toolbox here: https://github.com/obidam/pcm
 Catalogue PIGMA
Catalogue PIGMA