/Biological Environment/Bioinformatics
Type of resources
Available actions
Topics
Keywords
Contact for the resource
Provided by
Years
Representation types
-

DNA sequencing of Crassostrea gigas Pacific oyster spat infected in the wild with OsHV-1 virus in 4 French oyster basins (Marennes Oleron Bay, Arcachon bay, Rade de Brest and Thau lagoon).
-

The eleven collected wild strains of T. lutea were compared phenotypically, in particular with regard to their pigment and lipid profiles. The genome of each T. lutea strain was also sequenced to investigate the genetic structure and genome organisation of this species. Collected data were summarized in a genome browser to provide easy-to-use support for the scientific community (https://genomes-catalog.ifremer.fr). This provides an important resource- to understand, exploit and predict the biodiversity of this species.
-

Analysis of tuna stomach contents
-

In order to better characterize the genetic diversity of Cetaceans and especially the common Dolphin from the Bay of Biscay, sequences from the variable mitochondrial control region were obtained from water samples acquired close to groups of dolphins.
-

Successive infections with Vibrio harveyi were conducted in two populations of the European abalone in order to examine which genes may be involved in improved survival to the disease in the St. Malo population.
-

Individuals from 5 populations were kept in common garden conditions in order to examine acclimation and adaptation to temperature in the honeycomb worm. Worms were exposed to 5 temperature treatments, and collected for RNAseq analysis. Gene expression patterns were then examined.
-

ddRAD genotyping was used to evaluate population connectivity and putative loci under selection in honeycomb worm from 13 sites spanning its distribution in the Atlantic and Mediterranean coasts.
-

WGS of SARS-CoV-2 by Oxford Nanopore Technology from raw wastewater samples collected in France, 2020-2021
-
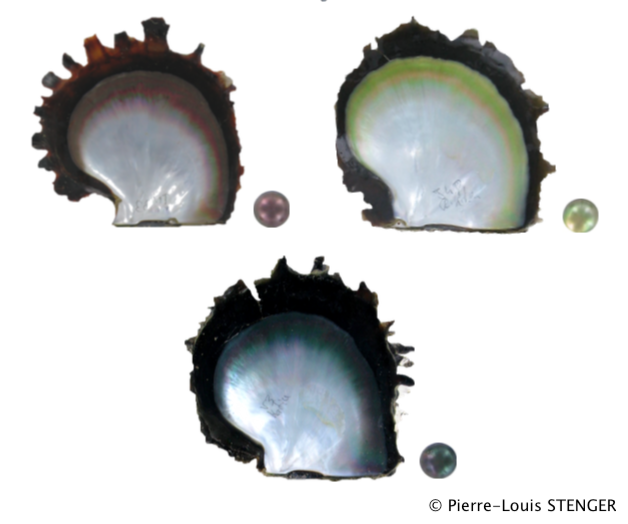

Whole genome pooled sequencing of individuals from 4 populations and 3 different color phenotype in order to uncover the genetic variants linked to color expression in the pearl oyster P. margaritifera.
-
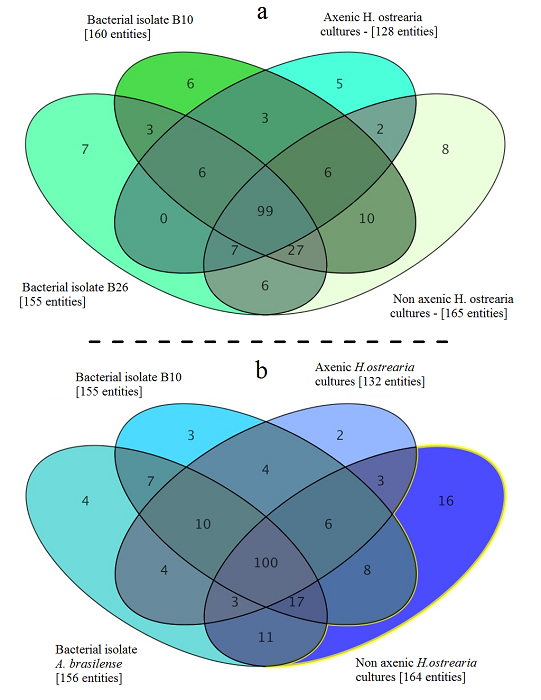
Metabolome of of the marine diatom Haslea ostrearia. Bacteria were isolated from Haslea ostrearia isolates cultivated in ES 1/3 medium in laboratory conditions over a 3-month period. These microalgal isolates were recovered from four sites on the French Atlantic coast: Bouin , La Barre-de-Monts (46.90 N; 2.11°W), Isle de Ré (46.22 N; 1.45°W), and La Tremblade (45.80 N; 1.15°W) . Data processing and statistical analysis of the metabolic profiles were performed on an LC/MS Metabolomics Discovery Workflow using Mass Profiler Professional Software and an Agilent 1290 Infinity II LC system coupled to an Agilent 6540 UHD Accurate-Mass QTOF hybrid mass spectrometer (Agilent Technologies, Waldbronn, Germany) equipped with a dual electrospray ionization (ESI) source. The full history (tools, parameters, input and output data files) is publicly available on http://dx.doi.org/10.12770/046e1e6a-864e-48a6-944b-d8613d67de0f
 Catalogue PIGMA
Catalogue PIGMA