/Sequences
Type of resources
Available actions
Topics
Keywords
Contact for the resource
Provided by
Years
Representation types
-
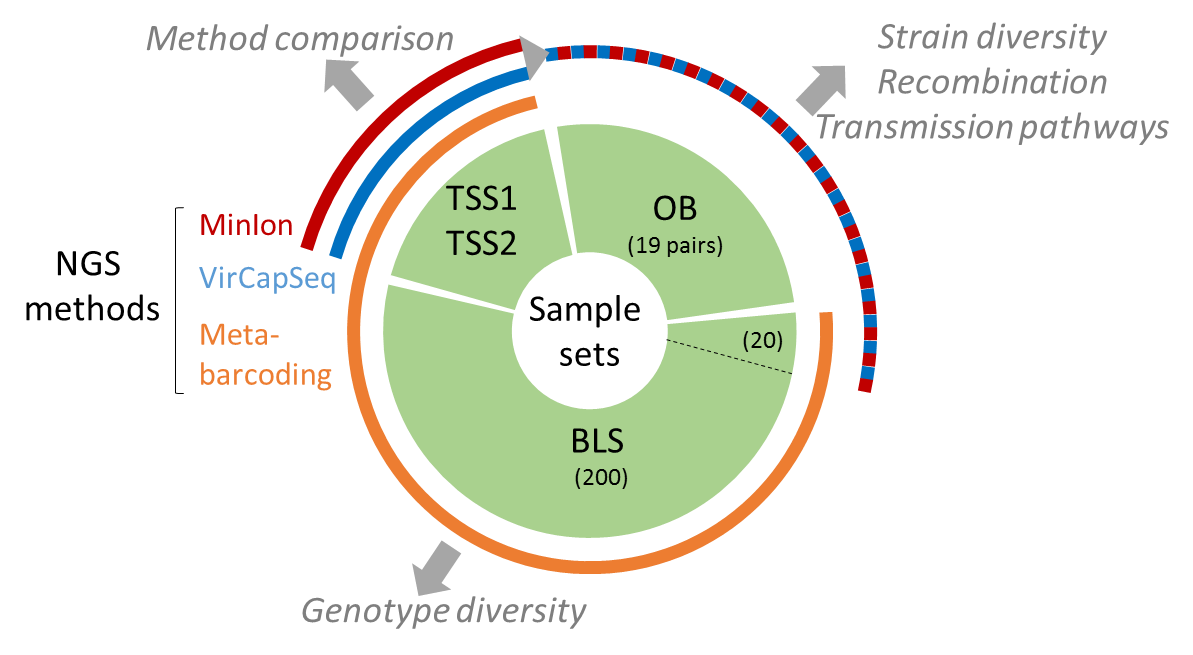
The present data set concerne metabarcoding raw reads produced using 4 different PCR targeting polymerase or capside coding region of the genoyupe I and II of norovirus. Test samples of norovirus with serial dilutions in pure water and after a bio-accumulation in oysters. Sequencing was made after VirCapSeq-VERT approach.
-

DNA sequencing of Crassostrea gigas Pacific oyster spat infected in the wild with OsHV-1 virus in 4 French oyster basins (Marennes Oleron Bay, Arcachon bay, Rade de Brest and Thau lagoon).
-

Dart Seq data gathered on Blue Shark in the framework of the PSTBS-IO project supported by funding from FAO, CSIRO Oceans and Atmosphere, AZTI Tecnalia, Institut de recherche pour le développement (IRD), and Research Institute for Tuna Fisheries (RITF) and financial assistance of the European Union (GCP/INT/233/EC – Population structure of IOTC species in the Indian Ocean), and POPSIZE project supported by FEAMP (2014-2020 UE N°508/2014), and Institut français de recherche pour l'Exploitation de la mer (Ifremer).
-

The project was designed to explore biological rhythms in the hydrothermal vent mussel Bathymodiolus azoricus. The experiment provides the first high-resolution temporal transcriptomes of an hydrothermal species, both in situ and in the laboratory.
-

Successive infections with Vibrio harveyi were conducted in two populations of the European abalone in order to examine which genes may be involved in improved survival to the disease in the St. Malo population.
-

In order to better characterize the fish biodiversity associated to the presence of cetaceans, especially the common Dolphin, from the Bay of Biscay, sequences from a 16S fragment were obtained from water samples acquired close to groups of dolphins.
-

DNA sequencing of Crassostrea gigas Pacific oyster spat experimentally infected with OsHV-1 virus from oyster basin of Marennes-Oleron
-

Here, our study aimed to first assess the influence of plastic on the bacterial community belonging to water, plastic and the microbiome of the giant clam and on the organism's physiology of this putative sentinel species. Our second objective was to identify bacteria whose abundance varies significantly with plastic concentration. Overall, this study will fill the gap towards a better understanding of the impact of plastic pollution on bacterial community assemblages in both inert and living environments.
-

In the framework of the PALDIAG project, environmental DNA from seawater was gathered in 2023 in Thau and Berre lagoons and studied with a 16S marker in order to detect the Veneroidae species.
-
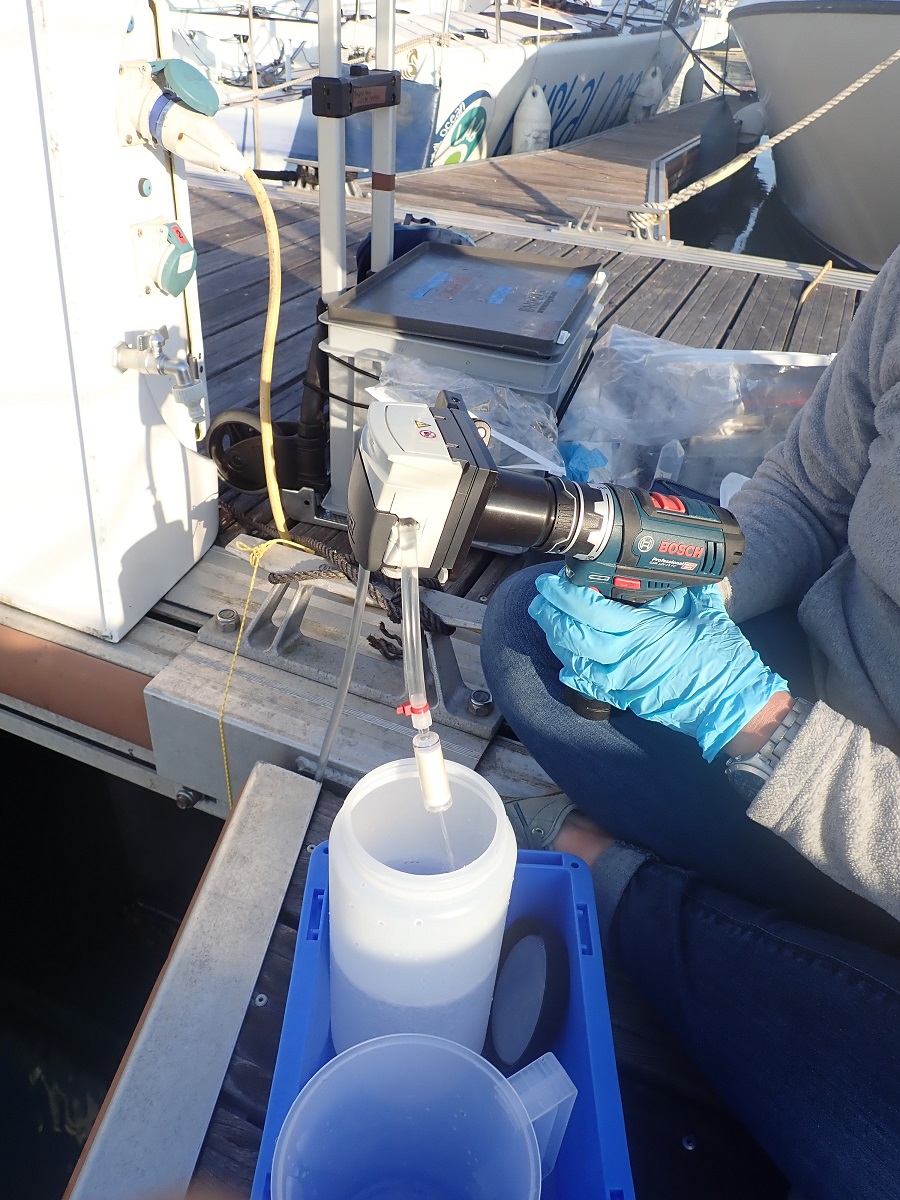

Metabarcoding data were produced based on samples gathered at Ifremer where the DNA was extracted; PCR libraries were built at Ifremer and Genseq; libraries were sequenced at Novogene. The data to download contain: 1/d emultiplexed raw data, 2/ metadata, and 3) Scripts to process data and taxonomically assign DNA sequences 4) Rmarkdown to analyze communities.
 Catalogue PIGMA
Catalogue PIGMA