2016
Type of resources
Available actions
Topics
Keywords
Contact for the resource
Provided by
Years
Formats
Representation types
Update frequencies
status
Scale
Resolution
-
Global Fishing Watch is working across the globe to provide governments and authorities with actionable reports and capacity building to help strengthen fisheries monitoring and compliance. Our global team of experts produce analyses to inform monitoring, control and surveillance of fisheries in five key areas: - Illegal, unreported and unregulated fishing - Transshipment - Port controls - Marine protected areas - Operation support Collaboration and information sharing are integral to achieving well-managed fisheries. By working with stakeholders and making analyses available to national, regional and intergovernmental partners, Global Fishing Watch is enabling fisheries agencies to make more informed and cost-efficient decisions. Topics: - Commercial fishing, Global Fishing Watch is harnessing innovative technology to turn transparent data into actionable information and drive tangible change in the way that fisheries are governed. - Transshipment, Through publicly sharing map visualisations and creating data and analysis tools, we seek to inform management and policy efforts and provide a more complete picture of transshipment at sea. - Marine protected areas, Global Fishing Watch is harnessing the data and technology revolution to support the effective design, management and monitoring of marine protected areas.
-

Présentation des entreprises, cartographie du risque et consignes en cas d'alerte pour les populations des communes de la Presqu'île d'Ambès. Plaquettes 4 ou 8 pages (avec cartographie)
-
Today's normative and regulatory requirements to assess the producible energy from wind rely on in situ measurements (mast with anemometric sensors), which are extremely costly to Implement offshore. However, proof should be provided that hindcast model results are highly reliable, in order to provide an equivalent assessment. Very high resolution models is also the key issue in decision making for a proper siting that is relaying on the consistency of all datasets provided in the assessment. In this tender the products of the FP7 MARINA project will be used. 10-year (2001-2010) highresolution atmospheric, wave, tidal and ocean current simulations will be used. The model outputs are at high resolution (0.05x0.05 degree horizontal resolution, 1-hour time resolution, 5-vertical levels at 10,40,80,120,180 m). The wave parameters are co-located with the meteorological output fields. Satellite altimetry data from ENVISAT and JASON satellites have been assimilated in the system. Other wind and wave satellite data sets will be also analyzed (Synthetic Aperture Radars-SAR for example). At the same co-located points the tidal and ocean current data together with bathymetry are available. For preselected points in the North Western Mediterranean (Spain-France-ltaly areas) directional wave spectra data have been saved and are available. From SKIRON meteorological model available parameters are: WIND SPEED (m/s), WIND DIRECTION (deg), AIR PRESSURE (hPa), AIR DENSITY (Kgr/m3), TEMPERATURE (K), MODEL SEAMASK From the wave model available parameters: SIGNIFICANT WAVE HEIGHT (m), MEAN WAVE DIRECTION (deg), WAVE MEAN PERIOD (s), PEAK WAVE PRERIOD (s), SWELL WAVE HEIGHT (m), MEAN SWELL PERIOD (s), MEAN DIRECTIONAL SPREAD, WINDSEA MEAN DIRECTIONAL SPREAD, SWELL MEAN DIRECTIONAL SPREAD, MAXIMUM WAVE HEIGHT (m)
-
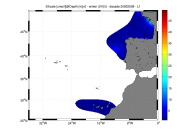

Moving 10-years analysis of Silicate at Northeast Atlantic Ocean for each season. - winter: January-March, - spring: April-June, - summer: July-September, - autumn: October-December Every year of the time dimension corresponds to the 10-year centred average of each season. Decades span - from 1963-1972 until 2005-2014 (winter) - from 1979-1988 until 2005-2014 (spring) - from 1990-1999 until 2005-2014 (summer) - from 1964-1973 until 2005-2014 (autumn) Observational data span from 1962 to 2014. Depth range (IODE standard depths): -3000.0, -2500.0, -2000.0, -1750, -1500.0, -1400.0, -1300.0, -1200.0, -1100.0, -1000.0, -900.0, -800.0, -700.0, -600.0, -500.0, -400.0, -300.0, -250.0, -200.0, -150.0, -125.0, -100.0, -75.0, -50.0,-40.0, -30.0, -20.0, -10.0, -5.0, -0.0 Data Sources: observational data from SeaDataNet/EMODNet Chemistry Data Network. Description of DIVA analysis: Geostatistical data analysis by DIVA (Data-Interpolating Variational Analysis) tool. GEBCO 1min topography is used for the contouring preparation. Analyzed filed masked using relative error threshold 0.3 and 0.5 DIVA settings. Signal to noise ratio and correlation length were optimized and filtered vertically and a seasonally-averaged profile was used. Logarithmic transformation applied to the data prior to the analysis. Background field: the data mean value is subtracted from the data. Detrending of data: no, Advection consraint applied: no. Units: umol/l
-
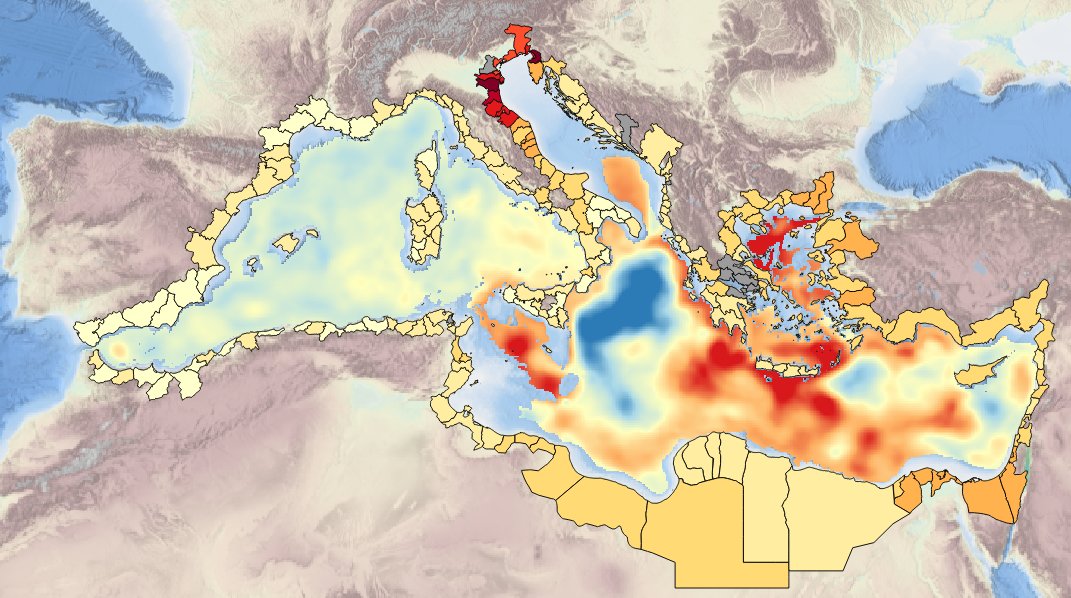
Specification of the desirable and recommended products attributes for generating spatial layers of sea mid-water and sea-bottom temperature for the last 10, 50 and 100 years for the Mediterranean basin and for each NUTS3 region along the coast.
-

Level 2 sub-skin Sea Surface Temperature derived from AVHRR on Metop, global and provided in full-resolution swath (1 km at nadir), in GHRSST compliant netCDF format. The satellite input data has successively come from Metop-A, Metop-B and Metop-C level 1 data processed at EUMETSAT. SST is retrieved from AVHRR infrared channels (3.7, 10.8 and 12.0 µm) using a multispectral algorithm and a cloud mask. Atmospheric profiles of water vapor and temperature from a numerical weather prediction model, Sea Surface Temperature from an analysis, together with a radiative transfer model, are used to correct the multispectral algorithm for regional and seasonal biases due to changing atmospheric conditions. The quality of the products is monitored regularly by daily comparison of the satellite estimates against buoy measurements.The product format is compliant with the GHRSST Data Specification (GDS) version 2. Users are advised to use data only with quality levels 3,4 and 5.
-
Specification of the desirable and recommended product attributes for generating time series of sea level trend for the last 10 years for the Mediterranean basin for each NUTS3 region along the coast.
-

Description of attributes for time series of sea level trend for the last 10 yrs for the Mediterranean basin and for each NUTS3 region along the coast.
-

The objective of this tender is to examine the current data collection, observation and data assembly programmes in the Meditterranean Sea, identify gaps and to evaluate how they can be optimised.
-
Data from FerryBoxes on ships of opportunity going on permanent routes are stored inside this database (ferrydata.hzg.de). Parameters are temperature, salinity, chlorophyll-a fluorescence, oxygen and different others. The data model is transect oriented. A data portal to access and visualise the data is also provided.
 Catalogue PIGMA
Catalogue PIGMA