CSV
Type of resources
Available actions
Topics
Keywords
Contact for the resource
Provided by
Years
Formats
Representation types
Update frequencies
status
Scale
Resolution
-
Global Fishing Watch is working across the globe to provide governments and authorities with actionable reports and capacity building to help strengthen fisheries monitoring and compliance. Our global team of experts produce analyses to inform monitoring, control and surveillance of fisheries in five key areas: - Illegal, unreported and unregulated fishing - Transshipment - Port controls - Marine protected areas - Operation support Collaboration and information sharing are integral to achieving well-managed fisheries. By working with stakeholders and making analyses available to national, regional and intergovernmental partners, Global Fishing Watch is enabling fisheries agencies to make more informed and cost-efficient decisions. Topics: - Commercial fishing, Global Fishing Watch is harnessing innovative technology to turn transparent data into actionable information and drive tangible change in the way that fisheries are governed. - Transshipment, Through publicly sharing map visualisations and creating data and analysis tools, we seek to inform management and policy efforts and provide a more complete picture of transshipment at sea. - Marine protected areas, Global Fishing Watch is harnessing the data and technology revolution to support the effective design, management and monitoring of marine protected areas.
-
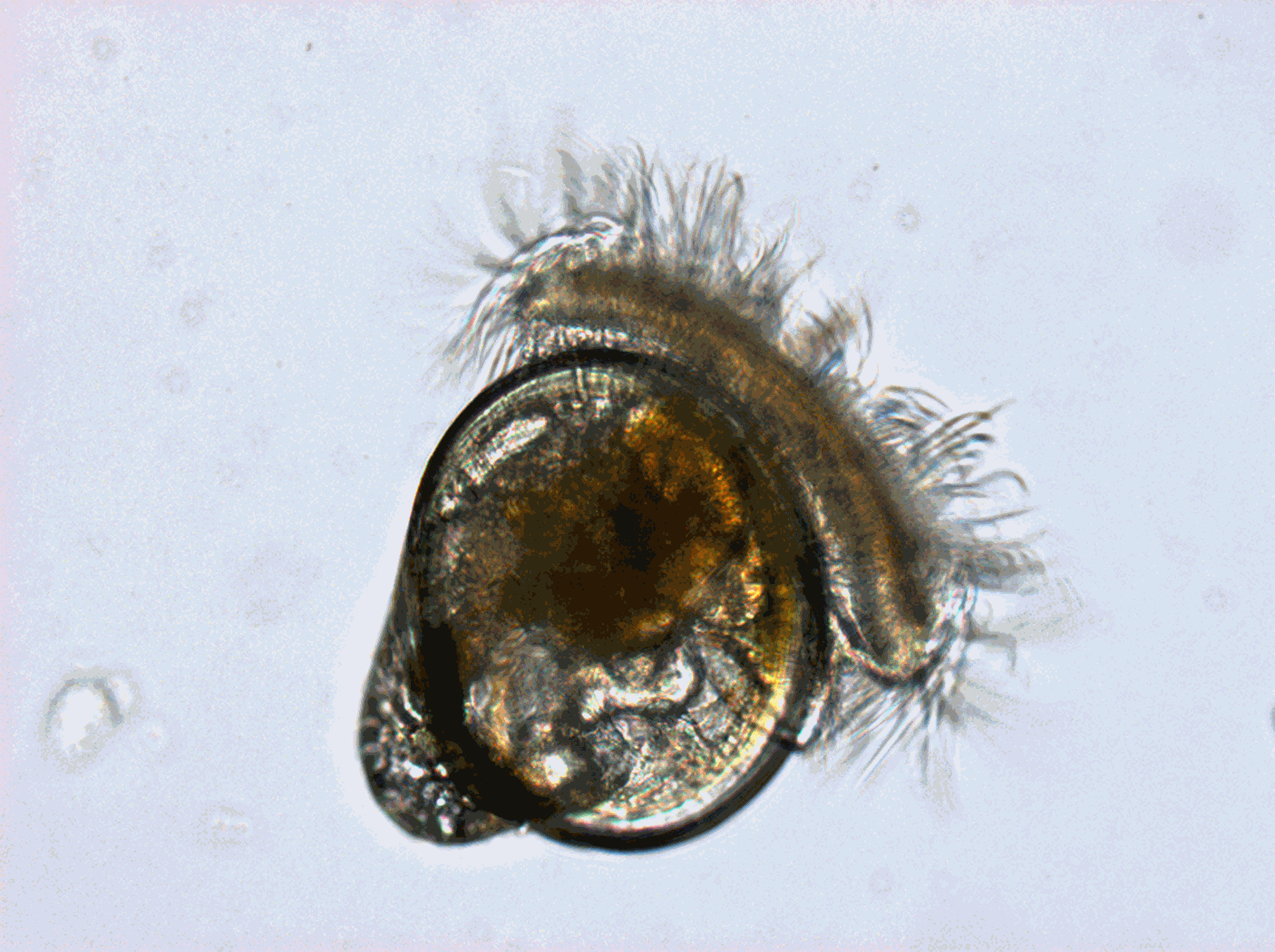
Worldwide, shellfish aquaculture and fisheries in coastal ecosystems represent crucial activities for human feeding. But these biological productions are under the pressure of climate variability and global change. Anticipating the biological processes affected by climate hazards remains a vital objective for species conservation strategies and human activities that rely on. Within marine species, filter feeders like oysters are real key species in coastal ecosystems due to their economic and societal value (fishing and aquaculture) but also due to their ecological importance. Indeed oysters populations in good health play the role of ecosystem engineers that can give many ecosystem services at several scales: building reef habitats that contribute to biodiversity, benthic-pelagic coupling and phytoplankton bloom control through water filtration, living shorelines against coastal erosion… The Pacific oyster, Crassostrea gigas (Thunberg, 1793), which is currently widespread worldwide, was introduced into the Atlantic European coasts at the end of the 19th century for shellfish culture purposes and becomes the main marine species farmed in France (around 100 000 tons) despite severe mortalities crisis. But in the same time and because of warming, natural oysters beds has spread significantly along the French coast and are supposed to have reach approximately 500 000 tons. In that context, Pacific oyster populations (natural and cultivated) in France are the subjects of many scientific projects. Among them, a specific long-term biological monitoring focuses on the reproduction of these populations at a national scale: the VELYGER national program. With more than 8 years of weekly data at many stations in France, this field-monitoring program offers a valuable dataset for studying processes underpinning reproduction cycle of this key-species in relation to environmental parameters, water quality and climate change. Database content: Larval concentration (number of individuals per 1.5 m3) monitored, since 2008, at several stations in six bays of the French coast (from south to north): Thau Lagoon and bays of Arcachon, Marennes Oléron, Bourgneuf, Vilaine and Brest (see map below). Methods used to monitor larval concentration: An important volume of seawater (1.5 m3) is pumped twice a week throughout the spawning season (june-september), at one meter below the surface at high tide (+/- 2h) in several sites within each VELYGER ecosystem. Water is filtered trough plankton net fitted with 40 µm mesh. After a proper rinsing of the net, the retained material is transferred into a polyethylene bottle (1 liter) and fixed with alcohol. At laboratory, sample is then gently filtered and rinse again and transferred into eprouvette. Two sub-samples of 1 mL are then taken using a pipette and examined on a graticule slide for microscope. The microscopic examination is made with a conventional binocular optical microscope with micrometer stage at a magnification of 10 X (or above). During the counting, a special care is necessary as larvae of other bivalves are also collected and confusion is possible. Larvae of C. gigas are also classified into four stage of development: - Stage I = D-shaped straight hinge larvae (shell length <105 µm) - Stage II = Early umbo evolved larvae (shell length between 105 and 150 µm) - Stage III = Medium umbo larvae (shell length between 150 and 235 µm) - Stage IV*= Large umbo eyed pediveliger larvae (shell length > 235 µm) * Larvae that are very closed to settle are sometimes identified into a separated 5th stage, but generally this stage is included in stage IV. Illustrations: Location of the different Velyger sites along the French coast. From south to north: Thau Lagoon and bays of Arcachon, Marennes Oléron, Bourgneuf, Vilaine and Brest. Legend: Pacific Oyster Larvae (left side) and Natural oyster bed (right side). Photos : © S. Pouvreau/Ifremer
-

To deliver the best Argo data to users in the simplest way, No QC flags; No data mode; No manuals - Just straight forward good data The Argo program provides an unprecedented volume of oceanographic data, yet its operational complexity — involving multiple data modes, quality control flags, and metadata conventions — often hinders its direct usage. The EasyOneArgo initiative addresses this challenge by delivering simplified, high-quality subsets of Argo data, specifically designed to streamline user access and integration. We introduce two core products: EasyOneArgoTS, a curated selection of temperature-salinity profiles filtered by strict quality criteria and optimized across real-time, adjusted, and delayed modes; and EasyOneArgoTSLite, its vertically interpolated counterpart standardized over 102 pressure levels. Each profile is packaged as a standalone CSV file with structured metadata, and indexed for seamless retrieval. Visual comparisons reveal clear advantages in usability and consistency, notably between raw and interpolated datasets. The approach is being extended to biogeochemical variables via EasyOneArgoBGC and EasyOneArgoBGCLite, currently under development. EasyOneArgo products are publicly available through monthly FAIR-compliant releases and invite community feedback for continued refinement. This work represents a user-centric shift in Argo data delivery: no flags, no manuals — just clean, structured ocean data ready for immediate scientific application.
-

Three saltmarshes, Aiguillon, Brouage, Fier d'Ars, located in the Pertuis-Charentais Sea along the south-west coast of France, were studied to evaluate their sediment and mass accumulation rates (SAR; MAR) based on 210Pb and 137Cs profiles in sediments. Coastal saltmarshes play indeed an essential role in providing services such as coastal protection and supporting biodiversity. Saltmarshes are also critical environments for the accumulation of sedimentary organic carbon (blue carbon). However, the number of studies on saltmarshes remains underrepresented compared to studies on mangroves and seagrass. This work is a contribution to the effort to document sediment and mass accumulation rates of saltmarshes.A total of 16 1m sediment cores were collected in the three saltmarshes (Aiguillon, Brouage, Fier d'Ars) in 2021 and 2022 using an Eijkelkamp stainless steel peat sampler. Each sediment core was sampled every 1 cm from the top to the bottom of the core. The sediment layers were used to determine dry bulk density and selected radioisotope activities (210Pb, 226Ra, 137Cs, 228Th, 137Cs). Combining excess 210Pb and 137Cs has allowed to establish a reliable chronology of sediment deposition on a multidecadal timescale.
-

The aim of this work was to document the seasonal and inter-annual dynamic of dissolved oxygen and ancillary data (T, S, Chl-a, turbidity, pH) along a cross-shelf transect off the Gironde estuary. This work has been motivated by recent simulations that suggest the occurrence of seasonal bottom deoxygenations in this River-dominated Ocean Margin (Riomar); but unfortunately there were no data sets to test this hypothesis until now. Profiles of temperature, salinity and dissolved oxygen were performed in the water column of the West Gironde Mud Patch off the Gironde estuary (from 45°46.383’N – 1°28.925’W to 45°35.524’N - 1°50.689’W) during seven cruises on the R/V Côte de la Manche (doi: 10.18142/284 ; 10.17600/18000861) between 2016 and 2021 (October 2016, August 2017, January 2018, April 2018, July 2019, April 2021, October 2021). Turbidity was measured in January and April 2018, July 2019 and October 2021, Chl-a in October 2016, August 2017, January 2018, April 2018 and July 2019 and pH in October 2021. This dataset had permitted to validate the occurrence of bottom deoxygenations when the water column is stratified.
-
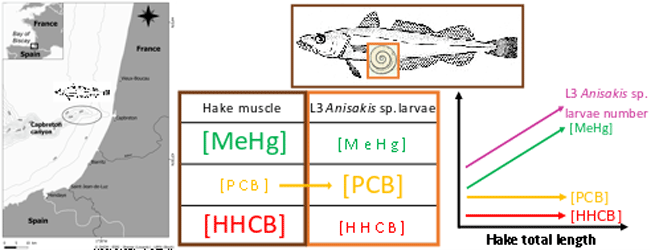
This dataset gathers isotopic ratios (carbon and nitrogen) and concentrations of both priority (mercury species and polychlorinated biphenyls congeners) and emerging (musks and sunscreens) micropollutants measured in a host-parasite couple (hake Merluccius merluccius muscle and in its parasite Anisakis sp) from the south of Bay of Biscay in 2018. In addition, the hake infection degree measured as the number of Anisakis sp. larvae was added for each hake collected.
-
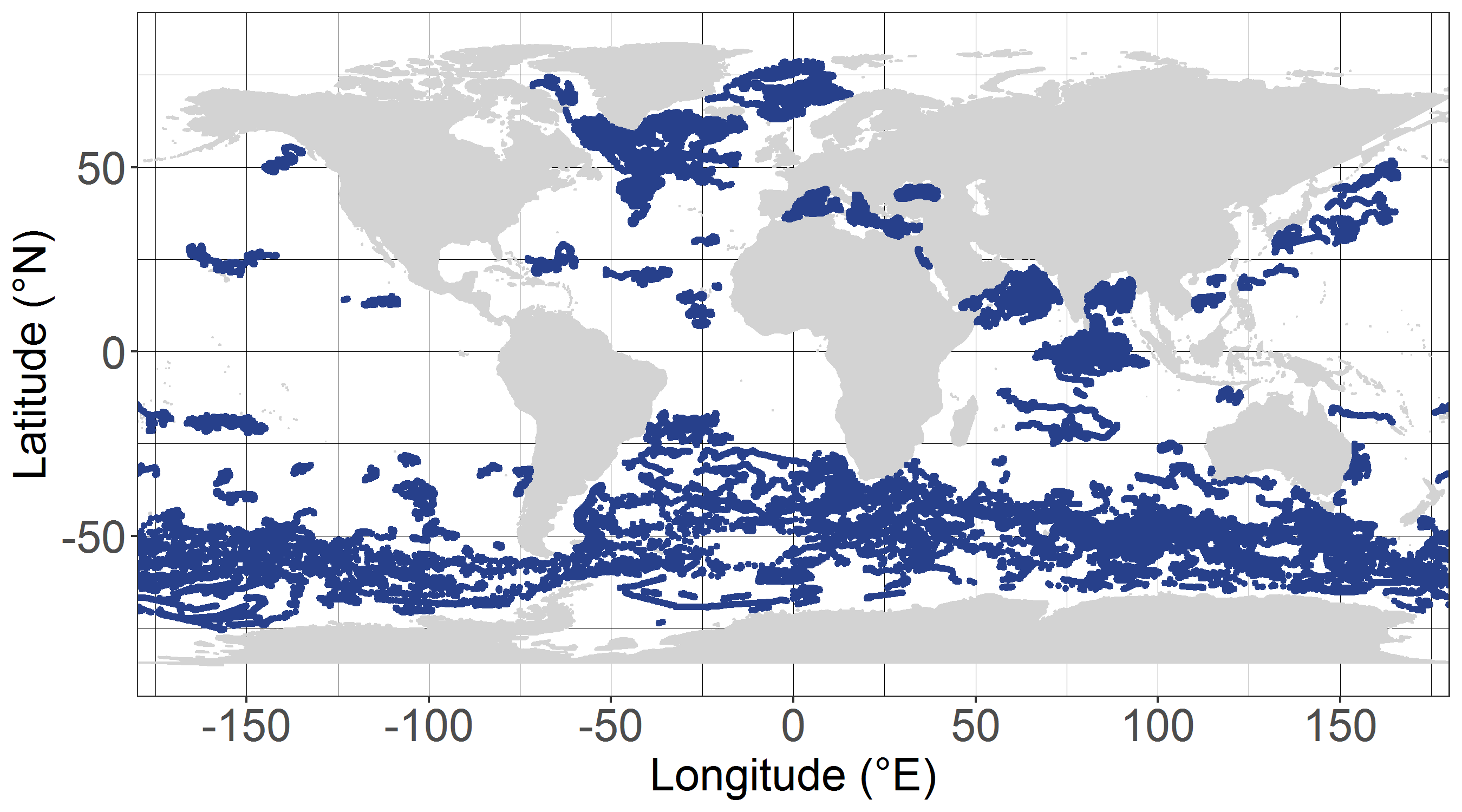
The dataset dcm_dtb.txt contains bio-optical measurements and environmental parameters associated with Deep Chlorophyll Maxima (DCM) acquired by BGC-Argo profiling floats. For each BGC-Argo profile the data files includes the World Meteorological Organization (WMO) and profile numbers, the Data Assembly Center (DAC), the geographical position (LON and LAT), the date of the profile in Julian Day (JULD) and in YYYY-MM-DD format; the region of the profile (REGION, acronyms detailed in the region.txt file), the DCM zonal attribution (ZONE, acronyms detailed in the zone.txt file), the vertical resolution of measurements of the concentration of the chlorophyll a [Chla] and of the backscattering coefficient (bbp) within the 250 first meters, the Mixed Layer Depth (MLD, m), the qualification of the vertical profile (DCM_TYPE) as Deep Biomass Maximum (3), Deep photoAcclimation Maximum (2), or presenting no DCM (1); the depth of the DCM (DCM_DEPTH); the chlorophyll a concentration (CHLA_DCM, mg chla m-3 ) the backscattering coefficient (BBP_DCM, m-1), and the Brunt-Vaisala frequency (N2_DCM) at the DCM depth; the nitracline depth (NCLINE_DEPTH, m) and steepness (NCLINE_STEEP, µmol NO3 m-3 m-1), the mean nitrate concentration within the Mixed Layer (NO3_MEAN_MLD, µmol NO3 m-3), the mean daily Photosynthetically Available Radiation in the Mixed Layer (MEAN_IPAR_MLD, E m -1 d -1), the daily Photosynthetically Available Radiation at the nitracline depth (IPAR_NCLINE, E m-2 d-1); and the [Chla] measured by satellite (CHLA_SAT, mg chla m-3). The dataset shape_NASTG_ASEW.txt contains the seasonal median, the first and third quartiles of the [Chla] and of the bbp profiles for the North Atlantic Subtropical Gyre and Atlantic SubEquatorial Waters regions. The dataset climato_NASTG_ASEW.txt contains the monthly mean and standard deviations of the DCM depth (DCM_depth), the isolume depth of daily Photosynthetically Available Radiation of 20 E m-2 d-1 (iPAR_20), the nitracline depth, and the Mixed Layer Depth (MLD) for the profiles within the North Atlantic Subtropical Gyre and Atlantic SubEquatorial Waters regions. The qualification and processing of the BGC-Argo profiles, as well as the DCM detection (DCM_TYPE) and the estimation of the environmental parameters, were applied as described from Cornec, M., Claustre, H., Mignot, A., Guidi, L., Lacour, L., Poteau, A., D’Ortenzio, F.,Gentili, B., Schmechtig, C., (to be updated.) Deep Chlorophyll Maxima in the global ocean: occurrences, drivers and characteristics. Global Biogeochemical Cycles, to be updated The [Chla] satellite variable was obtained by the match of each BGC-Argo profile with a L3S [Chla] product from the Ocean Colour-Climate Change Initiative v4.0 database merging observations from MERIS, MODIS, VIIRS and SeaWiFs, at a monthly and 4x4-km-pixel resolution, up to December 31, 2019 (ftp://oc-cci-data:ELaiWai8ae@oceancolour.org/occci-v4.2/).
-
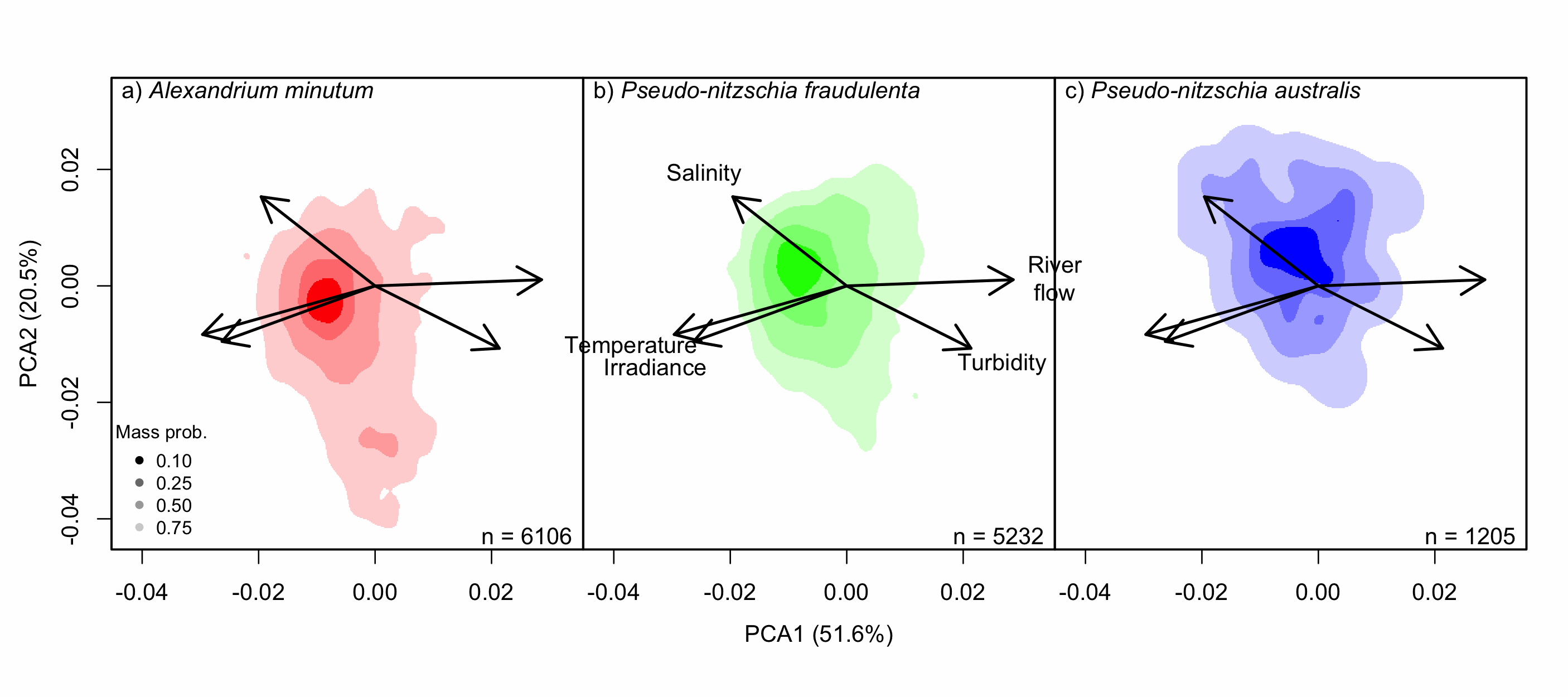
Understanding the spatial and temporal preferences of toxic phytoplankton species is of paramount importance in managing and predicting harmful events in aquatic ecosystems. In this study we address the realised niche of the species Alexandrium minutum, Pseudo-nitzschia fraudulenta and P. australis. We used them to highlight distribution patterns at different scales and determine possible drivers. To achieve this, we have developed original procedures coupling niche theory and habitat suitability modelling using abundance data in four consecutive steps: 1) Estimate the realised niche applying kernel functions. 2) Assess differences between the species’ niche as a whole and at the local level. 3) Develop habitat and temporal suitability models using niche overlap procedures. 4) Explore species temporal and spatial distributions to highlight possible drivers. Data used are species abundance and environmental variables collected over 27 years (1988-2014) and include 139 coastal water sampling sites along the French Atlantic coast. Results show that A. minutum and P. australis niches are very different, although both species have preference for warmer months. They both respond to decadal summer NAO but in the opposite way. P. fraudulenta realised niche lies in between the two other species niches. It also prefers warmer months but does not respond to decadal summer NAO. The Brittany peninsula is now classified as an area of prevalence for the three species. The methodology used here will allow to anticipate species distribution in the event of future environmental challenges resulting from climate change scenarios.
-

As part of the marine water quality monitoring of the “Pertuis” and the “baie de l’Aiguillon” (France), commissioned by the OFB and carried out by setec énergie environnement, three monitoring stations were installed. Two of them were set up at the mouths of the Charente and Seudre rivers on February 6 and 27, 2019, respectively, while a third was deployed in the Bay of Aiguillon on March 24, 2021. The dataset presented here concerns the station installed in the Charente estuary. Measurements are organized into .csv files, with one file per year. Data is collected using a SAMBAT multiparameter probe, which records the following parameters: - Temperature (-5 to 35 °C) - Conductivity (0 to 10 mS/cm) - Pressure (0 to 10 m) - Turbidity (0 to 300 NTU) - Dissolved Oxygen (0 to 20 mg/L & 0 to 200 %) - Fluorescence (0 to 50 µg/l) - PH (0/14)
-
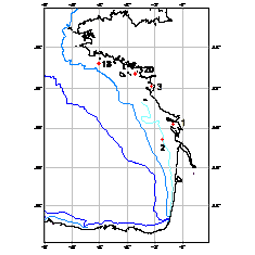
210Pb, 226Ra and 137Cs were measured by non-destructive gamma spectrometry on marine sediment cores, collected during RIKEAU 2002 cruise on board r/v Thalia, on the shelf of the Bay of Biscay
 Catalogue PIGMA
Catalogue PIGMA