2021
Type of resources
Available actions
Topics
Keywords
Contact for the resource
Provided by
Years
Formats
Representation types
Update frequencies
status
Scale
Resolution
-
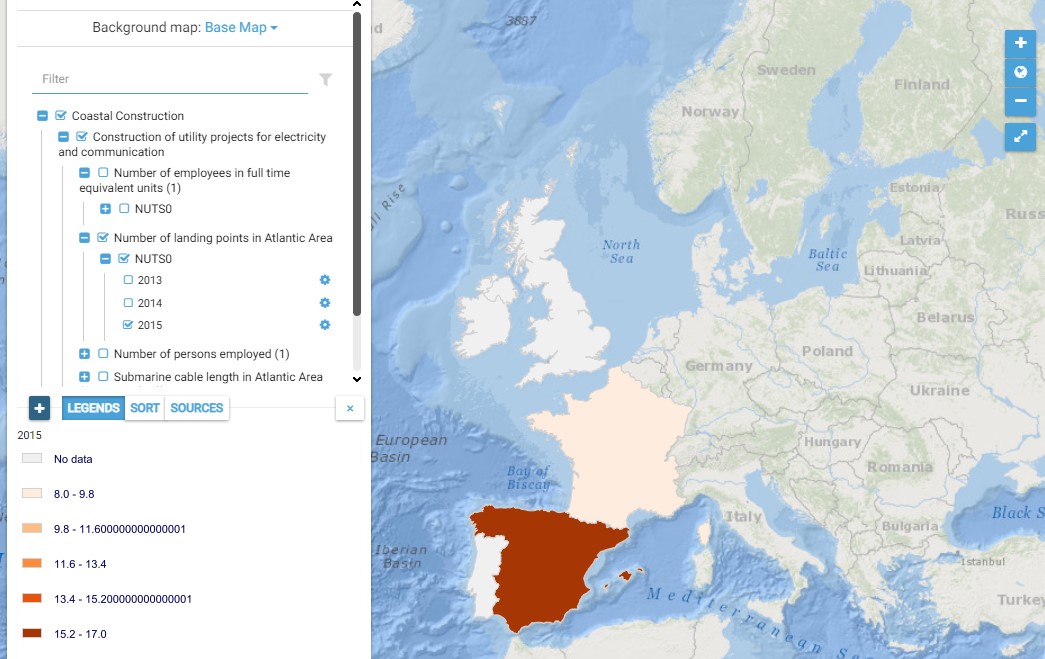
This map presents all layers corresponding to "Construction of utility projects for electricity and communication" activities in the Atlantic area. For more information about this NACE code : https://ec.europa.eu/eurostat/ramon/nomenclatures/index.cfm?TargetUrl=DSP_NOM_DTL_VIEW&StrNom=NACE_REV2&StrLanguageCode=EN&IntPcKey=18508154&IntKey=18508214&StrLayoutCode=HIERARCHIC&IntCurrentPage=1 Indicators collected are : Number of persons employed in cable and pipe laying and maintenance activities in the Atlantic area per country Submarine pipe length in Atlantic Area per country (P16) Number of landing points in Atlantic Area per country
-
NOAA STAR produces two lines of gridded 0.02deg super-collated L3S LEO datasets from Low Earth Orbiting (LEO) satellites, one from the NOAA afternoon JPSS (L3S_LEO_PM) and the other from the EUMETSAT mid-morning Metop-FG (L3S_LEO_AM). The L3S_LEO_PM is derived from JPSS satellites (in v2.80, NPP and N20) with VIIRS sensor onboard (0.75km/nadir). The L3S_LEO_PM dataset is produced by aggregating L3U datasets from two JPSS satellites ( https://doi.org/10.5067/GHVRS-3UO28 and https://doi.org/10.5067/GHV20-3UO28 ) and covers from Feb 2012-present. The L3S-LEO-PM data are reported in two files per 24hr interval, one daytime and one nighttime (nominal JPSS local equator crossing times around 01:30/13:30). Data is in NetCDF4 format, compliant with the GHRSST Data Specification version 2 (GDS2). The Near-Real Time (NRT) L3S-LEO data are archived at PO.DAAC with approximately 6 hours latency and then replaced by the Delayed Mode files about 2 months later, with identical file names. In addition to SST, the L3S-LEO files report the location and intensity of thermal fronts. The NRT/DM data are seamlessly stitched with the full-mission Reanalysis (RAN). The ACSPO L3S products are monitored and validated against in situ data in the NOAA iQuam system ( https://www.star.nesdis.noaa.gov/socd/sst/iquam ) in the NOAA SQUAM system ( https://www.star.nesdis.noaa.gov/socd/sst/squam ). Quality of SST imagery, clear-sky mask and thermal fronts is evaluated in the NOAA ARMS system ( https://www.star.nesdis.noaa.gov/socd/sst/arms ). NOAA plans to include data from other afternoon platforms and sensors, such as N21 and Aqua MODIS, into the future releases of the L3S_LEO_PM.
-
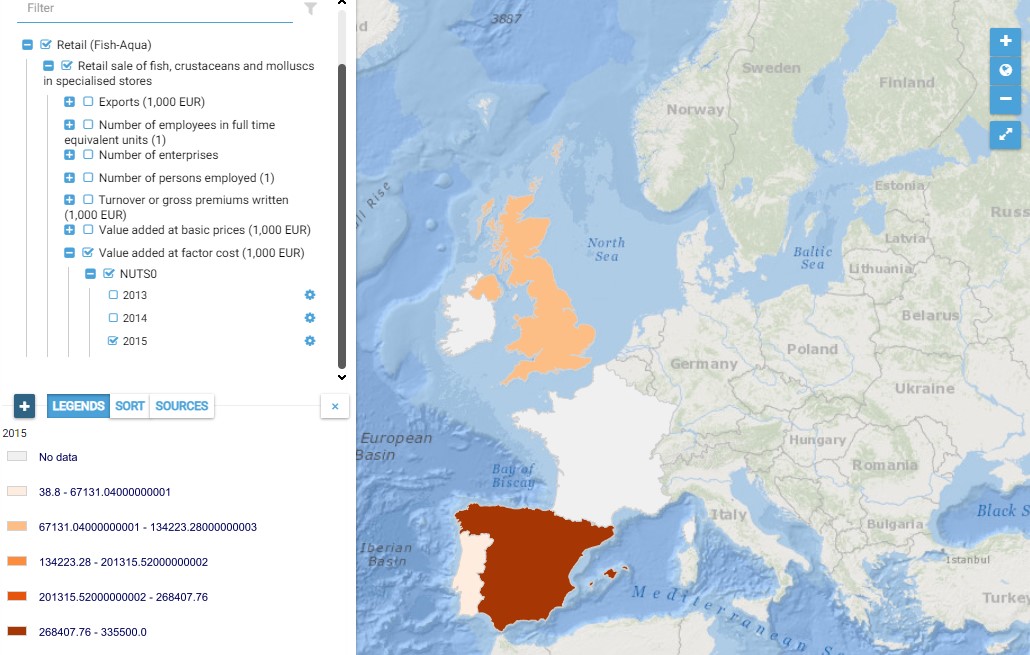
This map presents all layers corresponding to "Retail sale of fish, crustaceans and molluscs in specialised stores" activities in the Atlantic area. For more information about this NACE code : https://ec.europa.eu/eurostat/ramon/nomenclatures/index.cfm?TargetUrl=DSP_NOM_DTL_VIEW&StrNom=NACE_REV2&StrLanguageCode=EN&IntPcKey=18511064&IntKey=18511154&StrLayoutCode=HIERARCHIC&IntCurrentPage=1 Indicators collected are : Business indicators per country
-
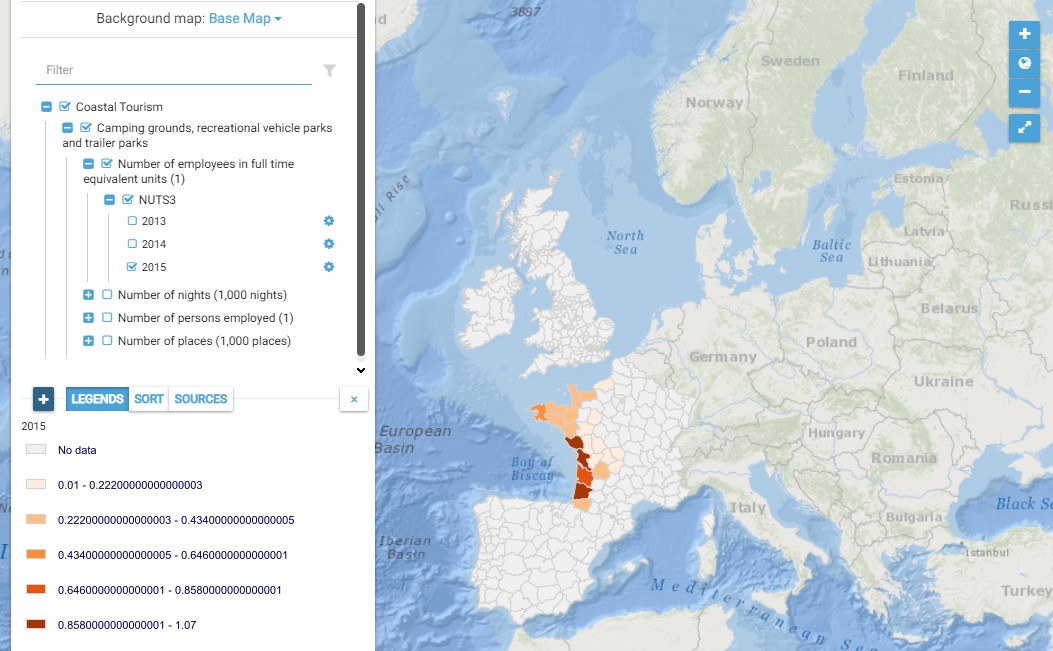
This map presents all layers corresponding to "Camping grounds, recreational vehicle parks and trailer parks" activities in the Atlantic area. For more information about this NACE code : https://ec.europa.eu/eurostat/ramon/nomenclatures/index.cfm?TargetUrl=DSP_NOM_DTL_VIEW&StrNom=NACE_REV2&StrLanguageCode=EN&IntPcKey=18513734&IntKey=18513764&StrLayoutCode=HIERARCHIC&IntCurrentPage=1 Indicators collected are : Number of places per NUTS 3 unit of the Atlantic Area
-
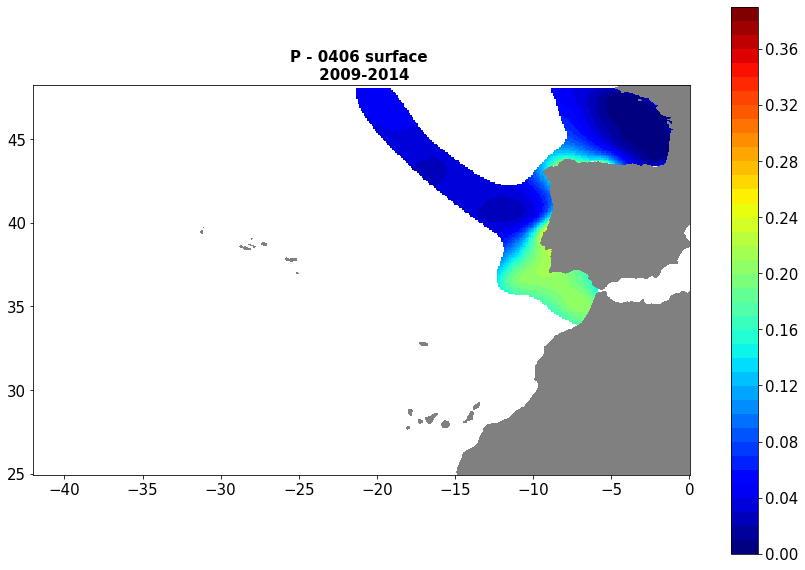
Moving 6-year analysis of Phosphate at Atlantic Sea for each season. - winter: January-March, - spring: April-June, - summer: July-September, - autumn: October-December Every year of the time dimension corresponds to the 6-year centred average of each season. 6-year periods span - from 1967-1972 until 2015-2020 (winter), - from 1960-1965 until 2015-2020 (spring), - from 1968-1973 until 2015-2020 (summer), - from 1961-1966 until 2015-2020 (autumn). Observational data span from 1960 to 2020. Depth range (IODE standard depths): -2000.0, -1750, -1500.0, -1400.0, -1300.0, -1200.0, -1100.0, -1000.0, -900.0, -800.0, -700.0, -600.0, -500.0, -400.0, -300.0, -250.0, -200.0, -150.0, -125.0, -100.0, -75.0, -50.0,-40.0, -30.0, -20.0, -10.0, -5.0, -0.0 Data Sources: observational data from SeaDataNet/EMODNet Chemistry Data Network. Description of DIVA analysis: Geostatistical data analysis by DIVA (Data-Interpolating Variational Analysis) tool. GEBCO 1min topography is used for the contouring preparation. Analyzed filed masked using relative error threshold 0.3 and 0.5 DIVA settings. Correlation length was optimized and filtered vertically and a seasonally-averaged profile was used. Signal to noise ratio was fixed to 1. Logarithmic transformation applied to the data prior to the analysis. Background field: the data mean value is subtracted from the data. Detrending of data: no, Advection constraint applied: no. Units: umol/l
-
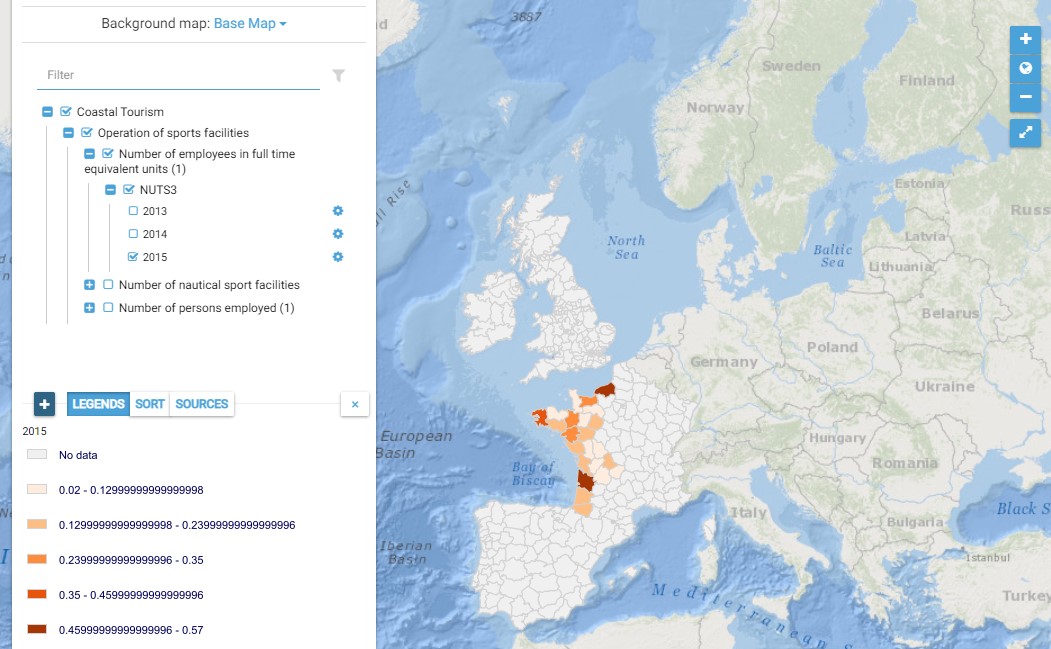
This map presents all layers corresponding to "Operation of sports facilities" activities in the Atlantic area. For more information about this NACE code : https://ec.europa.eu/eurostat/ramon/nomenclatures/index.cfm?TargetUrl=DSP_NOM_DTL_VIEW&StrNom=NACE_REV2&StrLanguageCode=EN&IntPcKey=18522104&IntKey=18522134&StrLayoutCode=HIERARCHIC&IntCurrentPage=1 Indicators collected are : - Number of persons employed and number of employees in full time equivalent units per NUTS 3 unit of the Atlantic Area - Number of nautical sport facilities per NUTS3 unit of the Atlantic Area
-
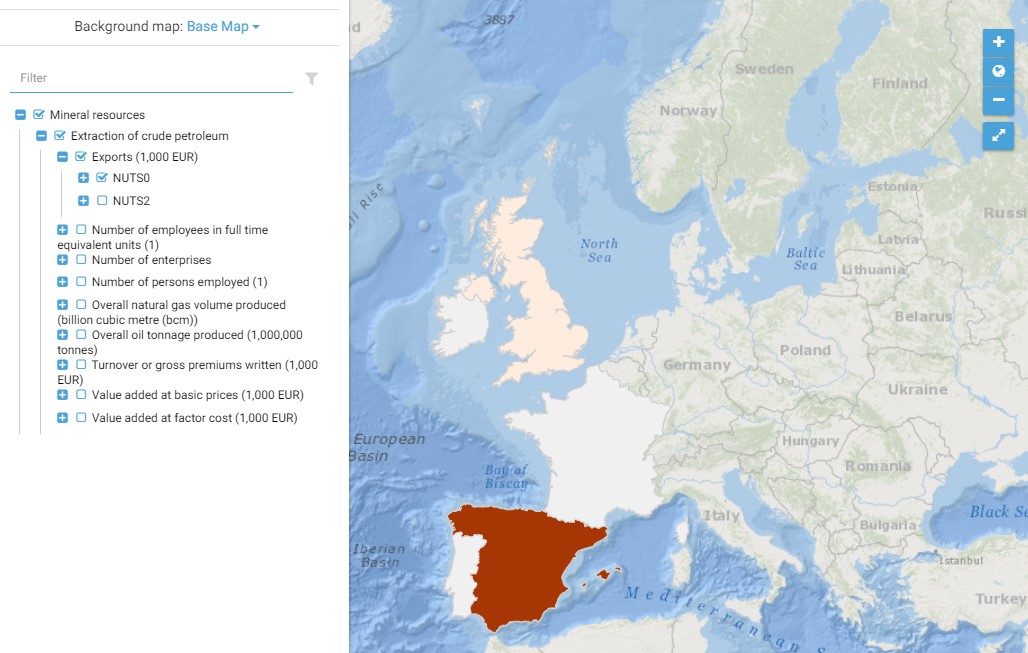
This map presents all layers corresponding to "Extraction of crude petroleum" activities in the Atlantic area. For more information about this NACE code : https://ec.europa.eu/eurostat/ramon/nomenclatures/index.cfm?TargetUrl=DSP_NOM_DTL_VIEW&StrNom=NACE_REV2&StrLanguageCode=EN&IntPcKey=18495614&IntKey=18495644&StrLayoutCode=HIERARCHIC&IntCurrentPage=1 Indicators collected are : Business indicators per country Number of persons employed on Atlantic pits and rigs Overall oil tonnage produced Overall volume produced
-
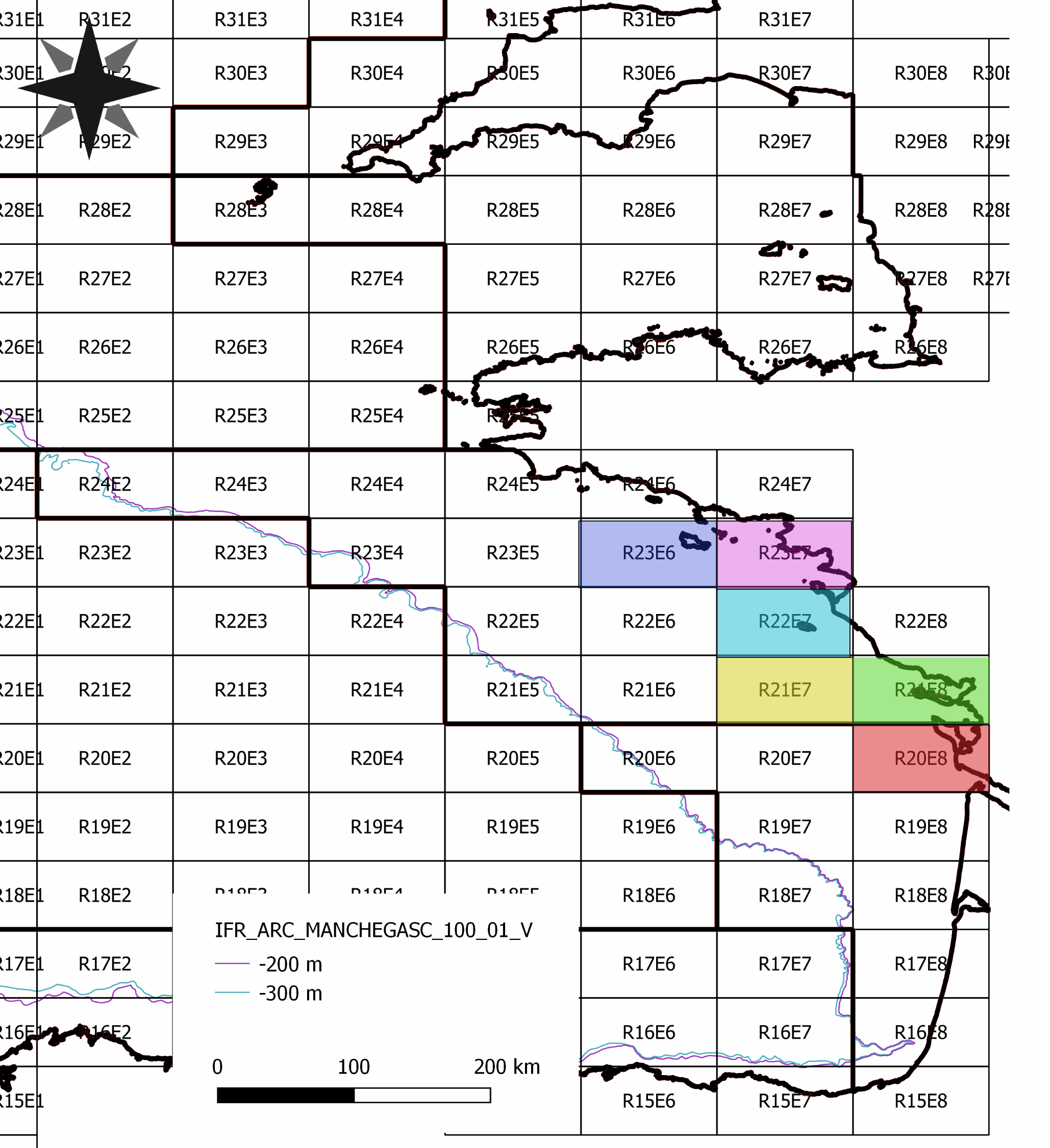
############# # Data description # ############# This dataset have been constructed and used for scientific purpose, available in the paper "Detecting the effects of inter-annual and seasonal changes of environmental factors on the the striped red mullet population in the Bay of Biscay" authored by Kermorvant C., Caill-Milly N., Sous D., Paradinas I., Lissardy M. and Liquet B. and published in Journal of Sea Research. This file is an extraction from the SACROIS fisheries database created by Ifremer (for more information see https://sextant.ifremer.fr/record/3e177f76-96b0-42e2-8007-62210767dc07/) and from the Copernicus database. Biochemestry comes from the product GLOBAL_ANALYSIS_FORECAST_BIO_001_028 (https://resources.marine.copernicus.eu/?option=com_csw&view=details&product_id=GLOBAL_ANALYSIS_FORECAST_BIO_001_028). Temperature and salinity comes from GLOBAL_ANALYSIS_FORECAST_PHY_001_024 product (https://resources.marine.copernicus.eu/?option=com_csw&view=details&product_id=GLOBAL_ANALYSIS_FORECAST_PHY_001_024). As fisheries landing per unit of effort is only available per ICES rectangle and by month, environmental data have been aggregated accordingly. ############### # Colomns description # ############### rectangle - The 6 ICES statistical rectangles used in the study. time_m - Time in months, from the beginning to the end of the study. annee = year mois = month (from 1 to 12) Poids = Weight of red mullet landed valeur = Temps_peche = fishing time Nb_sequence = number of fishing sequences Moy / Med / Var / StD Quartil_1 / Quartil_3 / min / max / CV / IQR = statistical descriptors of landing by rectangle and by month log_cpue = log of Med colomn mean_surface_s = mean of surface salinity by month and by rectangle median_surface_s = median of surface salinity by month and by rectangle mean_surface_t = mean of surface temperature by month and by rectangle median_surface_t = median of surface temperature by month and by rectangle si / zeu /po4 / pyc / o2/ nppv / no3 and nh4 mean and median concentration by rectangle and by month pc3 / pc2 / pc1 - projections of previous biochemestry variables on the three first axes of a PCA
-
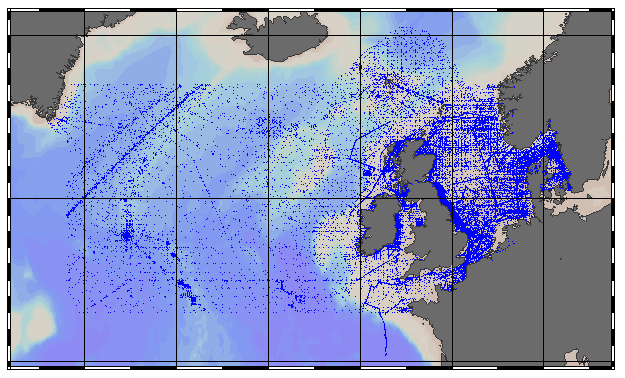
EMODnet Chemistry aims to provide access to marine chemistry data sets and derived data products concerning eutrophication, acidity and contaminants. The chemicals chosen reflect importance to the Marine Strategy Framework Directive (MSFD). ITS-90 water temperature and Water body salinity variables have been also included (as-is) to complete the Eutrophication and Acidity data. If you use these variables for calculations, please refer to SeaDataNet for having the quality flags: https://www.seadatanet.org/Products/Aggregated-datasets . This aggregated dataset contains all unrestricted EMODnet Chemistry data on Eutrophication and Acidity (14 parameters with quality flag indicators), and covers the North Sea with 587584 CDI records. Data were aggregated and quality controlled by 'Aarhus University, Department of Bioscience, Marine Ecology Roskilde' from Denmark. Regional datasets concerning eutrophication and acidity are automatically harvested and resulting collections are aggregated and quality controlled using ODV Software and following a common methodology for all Sea Regions ( https://doi.org/10.6092/9f75ad8a-ca32-4a72-bf69-167119b2cc12). When not present in original data, Water body nitrate plus nitrite was calculated by summing up the Nitrates and Nitrites. Same procedure was applied for Water body dissolved inorganic nitrogen (DIN) which was calculated by summing up the Nitrates, Nitrites and Ammonium. Quality flags for Water body dissolved inorganic nitrogen (DIN) should be disregarded since that currently they are not based on the original quality flags of nitrite, nitrate and ammonium. Parameter names are based on P35, EMODnet Chemistry aggregated parameter names vocabulary, which is available at: https://www.bodc.ac.uk/resources/vocabularies/vocabulary_search/P35/. Detailed documentation is available at: https://dx.doi.org/10.6092/4e85717a-a2c9-454d-ba0d-30b89f742713 Explore and extract data at: https://emodnet-chemistry.webodv.awi.de/eutrophication>NorthSea The aggregated dataset can be downloaded as ODV collection and spreadsheet, which is composed of metadata header followed by tab separated values. This spreadsheet can be imported to ODV Software for visualisation (More information can be found at: http://www.seadatanet.org/Standards-Software/Software/ODV). The original datasets can be searched and downloaded from EMODnet Chemistry Chemistry CDI Data and Discovery Access Service: https://emodnet-chemistry.maris.nl/search
-
Water body chlorophyll-a - Monthly Climatology for the European Seas for the period 1960-2020 on the domain from longitude -45.0 to 70.0 degrees East and latitude 24.0 to 83.0 degrees North. Data Sources: observational data from SeaDataNet/EMODnet Chemistry Data Network. Description of DIVA analysis: The computation was done with the DIVAnd (Data-Interpolating Variational Analysis in n dimensions), version 2.7.2, using GEBCO 30sec topography for the spatial connectivity of water masses. Horizontal correlation length and vertical correlation length vary spatially depending on the topography and domain. Depth range: 0.0, 5.0, 10.0, 15.0, 20.0, 25.0, 30.0, 35.0, 40.0, 45.0, 50.0, 55.0, 60.0, 65.0, 70.0, 75.0, 80.0, 85.0, 90.0, 95.0, 100.0, 125.0, 150.0, 175.0, 200.0, 225.0, 250.0, 275.0, 300.0, 325.0, 350.0, 375.0, 400.0, 425.0, 450.0, 475.0, 500.0, 550.0, 600.0, 650.0, 700.0, 750.0, 800.0, 850.0, 900.0, 950.0, 1000.0, 1050.0, 1100.0, 1150.0, 1200.0, 1250.0, 1300.0, 1350.0, 1400.0, 1450.0, 1500.0, 1550.0, 1600.0, 1650.0, 1700.0, 1750.0, 1800.0, 1850.0, 1900.0, 1950.0, 2000.0, 2100.0, 2200.0, 2300.0, 2400.0, 2500.0, 2600.0, 2700.0, 2800.0, 2900.0, 3000.0, 3100.0, 3200.0, 3300.0, 3400.0, 3500.0, 3600.0, 3700.0, 3800.0, 3900.0, 4000.0, 4100.0, 4200.0, 4300.0, 4400.0, 4500.0, 4600.0, 4700.0, 4800.0, 4900.0, 5000.0, 5100.0, 5200.0, 5300.0, 5400.0, 5500.0 m. Units: mg/m3. The horizontal resolution of the produced DIVAnd analysis is 0.25 degrees.
 Catalogue PIGMA
Catalogue PIGMA