2025
Type of resources
Available actions
Topics
Keywords
Contact for the resource
Provided by
Years
Formats
Representation types
Update frequencies
status
Service types
Scale
Resolution
-
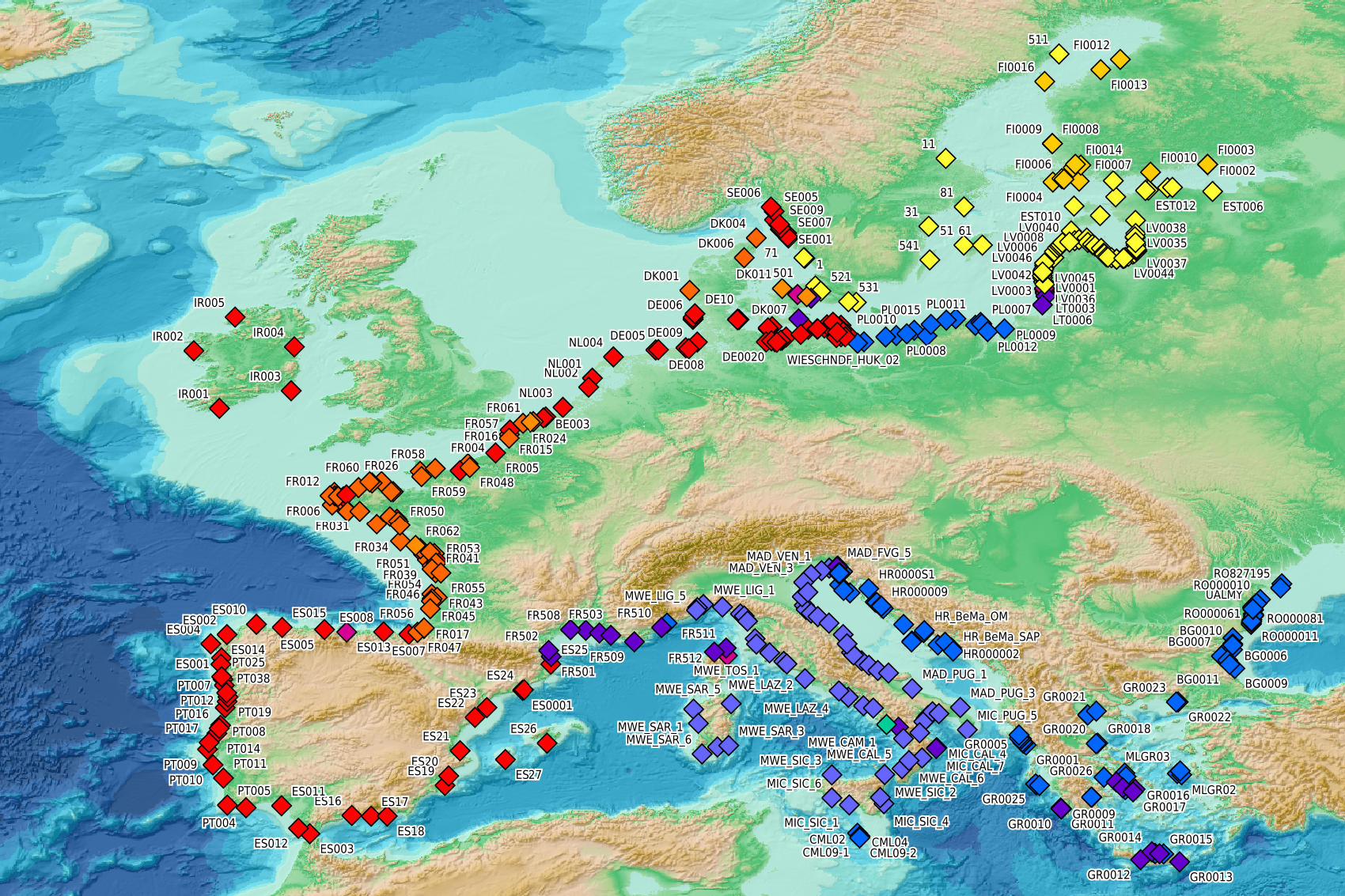
This visualization product displays beaches locations where the Marine Strategy Framework Directive (MSFD) monitoring protocol has been applied to collate data on macrolitter (> 2.5 cm). Reference lists associated with these protocols have been indicated with different colors in the map. EMODnet Chemistry included the collection of marine litter in its 3rd phase. Since the beginning of 2018, data of beach litter have been gathered and processed in the EMODnet Chemistry Marine Litter Database (MLDB). The harmonization of all the data has been the most challenging task considering the heterogeneity of the data sources, sampling protocols and reference lists used on a European scale. Preliminary processings were necessary to harmonize all the data: - Exclusion of OSPAR 1000 protocol: in order to follow the approach of OSPAR that it is not including these data anymore in the monitoring; - Selection of MSFD surveys only (exclusion of other monitoring, cleaning and research operations); - Exclusion of beaches without coordinates; - Some categories & some litter types like organic litter, small fragments (paraffin and wax; items > 2.5cm) and pollutants have been removed. This list was created using EU Marine Beach Litter Baselines, the European Threshold Value for Macro Litter on Coastlines and the Joint list of litter categories for marine macro-litter monitoring from JRC (these three documents are attached to this metadata). More information is available in the attached documents. Warning: the absence of data on the map does not necessarily mean that they do not exist, but that no information has been entered in the Marine Litter Database for this area.
-
Ensemble simulations of the ecosystem model Apecosm (https://apecosm.org) forced by the IPSL-CM6-LR climate model with the climate change scenario SSP5-8.5. The output files contain yearly mean biomass density for 3 communities (epipelagic, mesopelagic migratory and mesopelagic redidents) and 100 size classes (ranging from 0.12cm to 1.96m) The model grid file is also provided. Units are in J/m2 and can be converted in kg/m2 by dividing by 4e6. These outputs are associated with the "Assessing the time of emergence of marine ecosystems from global to local scales using IPSL-CM6A-LR/APECOSM climate-to-fish ensemble simulations" paper from the Earth's Future "Past and Future of Marine Ecosystems" Special Collection.
-
This delayed mode product designed for reanalysis purposes integrates the best available version of in situ data for ocean surface currents and current vertical profiles. It concerns three delayed time datasets dedicated to near-surface currents measurements coming from three platforms (Lagrangian surface drifters, High Frequency radars and Argo floats) and velocity profiles within the water column coming from the Acoustic Doppler Current Profiler (ADCP, vessel mounted only). The latest version of Copernicus surface and sub-surface water velocity product is also distributed from Copernicus Marine catalogue.
-

Moving 6-year analysis of Water body dissolved oxygen concentration in the Mediterranean Sea for each season: - winter: January-March, - spring: April-June, - summer: July-September, - autumn: October-December. Every year of the time dimension corresponds to the 6-year centered average of the season. 6-years periods span from 1970-1975 until 2019-2024. Description of DIVA analysis: The computation was done with the DIVAnd (Data-Interpolating Variational Analysis in n dimensions), version 2.7.12, using GEBCO 30sec topography for the spatial connectivity of water masses. The horizontal resolution of the produced DIVAnd maps grids is dx=dy=0.125 degrees (around 13.5km and 10.9km accordingly). The vertical resolution is 27 depth levels: [0.,5.,10.,20.,30.,50.,75.,100.,125.,150.,200.,250.,300.,400.,500.,600.,700.,800.,900.,1000.,1100.,1200.,1300.,1400.,1500.,1750.,2000.]. The horizontal correlation length is 200km. The vertical correlation length (in meters) was set twices the vertical resolution: [10.,10.,20.,20.,40.,50.,50.,50.,50.,100.,100.,100.,200.,200.,200.,200.,200.,200.,200.,200.,200.,200.,200.,200.,500.,500.,500.]. Duplicates check was performed using the following criteria for space and time: dlon=0.001deg., dlat=0.001deg., ddepth=1m, dtime=1hour, dvalue=0.1. The error variance (epsilon2) was set equal to 1 for profiles and 10 for time series to reduce the influence of close data near the coasts. An anamorphosis transformation was applied to the data (function DIVAnd.Anam.loglin) to avoid unrealistic negative values: threshold value=200. A background analysis field was used for all years (1970-2024) with correlation length equal to 600km and error variance (epsilon2) equal to 20. Quality control of the observations was applied using the interpolated field (QCMETHOD=3). Residuals (differences between the observations and the analysis (interpolated linearly to the location of the observations) were calculated. Observations with residuals outside the minimum and maximum values of the 99% quantile were discarded from the analysis. Originators of Italian data sets-List of contributors: - Brunetti Fabio (OGS) - Cardin Vanessa, Bensi Manuel doi:10.6092/36728450-4296-4e6a-967d-d5b6da55f306 - Cardin Vanessa, Bensi Manuel, Ursella Laura, Siena Giuseppe doi:10.6092/f8e6d18e-f877-4aa5-a983-a03b06ccb987 - Cataletto Bruno (OGS) - Cinzia Comici Cinzia (OGS) - Civitarese Giuseppe (OGS) - DeVittor Cinzia (OGS) - Giani Michele (OGS) - Kovacevic Vedrana (OGS) - Mosetti Renzo (OGS) - Solidoro C.,Beran A.,Cataletto B.,Celussi M.,Cibic T.,Comici C.,Del Negro P.,De Vittor C.,Minocci M.,Monti M.,Fabbro C.,Falconi C.,Franzo A.,Libralato S.,Lipizer M.,Negussanti J.S.,Russel H.,Valli G., doi:10.6092/e5518899-b914-43b0-8139-023718aa63f5 - Celio Massimo (ARPA FVG) - Malaguti Antonella (ENEA) - Fonda Umani Serena (UNITS) - Bignami Francesco (ISAC/CNR) - Boldrini Alfredo (ISMAR/CNR) - Marini Mauro (ISMAR/CNR) - Miserocchi Stefano (ISMAR/CNR) - Zaccone Renata (IAMC/CNR) - Lavezza, R., Dubroca, L. F. C., Ludicone, D., Kress, N., Herut, B., Civitarese, G., Cruzado, A., Lefèvre, D.,Souvermezoglou, E., Yilmaz, A., Tugrul, S., and Ribera d'Alcala, M.: Compilation of quality controlled nutrient profiles from the Mediterranean Sea, doi:10.1594/PANGAEA.771907, 2011.
-

Moving 6-year analysis of Water body chlorophyll-a in the Mediterranean Sea for each season: - winter: January-March, - spring: April-June, - summer: July-September, - autumn: October-December. Every year of the time dimension corresponds to the 6-year centered average of the season. 6-years periods span from 1990-1995 until 2019-2024. Description of DIVA analysis: The computation was done with the DIVAnd (Data-Interpolating Variational Analysis in n dimensions), version 2.7.12, using GEBCO 30sec topography for the spatial connectivity of water masses. The horizontal resolution of the produced DIVAnd maps grids is dx=dy=0.125 degrees (around 13.5km and 10.9km accordingly). The vertical resolution is 20 depth levels: [0.,5.,10.,20.,30.,50.,75.,100.,125.,150.,200.,250.,300.,400.,500.,600.,700.,800.,900.,1000.]. The horizontal correlation length is 200km. The vertical correlation length (in meters) was set twices the vertical resolution: [10.,10.,20.,20.,40.,50.,50.,50.,50.,100.,100.,100.,200.,200.,200.,200.,200.,200.,200.,200.]. Duplicates check was performed using the following criteria for space and time: dlon=0.001deg., dlat=0.001deg., ddepth=1m, dtime=1hour, dvalue=0.1. The error variance (epsilon2) was set equal to 1 for profiles and 10 for time series to reduce the influence of close data near the coasts. An anamorphosis transformation was applied to the data (function DIVAnd.Anam.loglin) to avoid unrealistic negative values: threshold value=200. A background analysis field was used for all years (1990-2024) with correlation length equal to 600km and error variance (epsilon2) equal to 20. Quality control of the observations was applied using the interpolated field (QCMETHOD=3). Residuals (differences between the observations and the analysis (interpolated linearly to the location of the observations) were calculated. Observations with residuals outside the minimum and maximum values of the 99% quantile were discarded from the analysis. Originators of Italian data sets-List of contributors: - Brunetti Fabio (OGS) - Cardin Vanessa, Bensi Manuel doi:10.6092/36728450-4296-4e6a-967d-d5b6da55f306 - Cardin Vanessa, Bensi Manuel, Ursella Laura, Siena Giuseppe doi:10.6092/f8e6d18e-f877-4aa5-a983-a03b06ccb987 - Cataletto Bruno (OGS) - Cinzia Comici Cinzia (OGS) - Civitarese Giuseppe (OGS) - DeVittor Cinzia (OGS) - Giani Michele (OGS) - Kovacevic Vedrana (OGS) - Mosetti Renzo (OGS) - Solidoro C.,Beran A.,Cataletto B.,Celussi M.,Cibic T.,Comici C.,Del Negro P.,De Vittor C.,Minocci M.,Monti M.,Fabbro C.,Falconi C.,Franzo A.,Libralato S.,Lipizer M.,Negussanti J.S.,Russel H.,Valli G., doi:10.6092/e5518899-b914-43b0-8139-023718aa63f5 - Celio Massimo (ARPA FVG) - Malaguti Antonella (ENEA) - Fonda Umani Serena (UNITS) - Bignami Francesco (ISAC/CNR) - Boldrini Alfredo (ISMAR/CNR) - Marini Mauro (ISMAR/CNR) - Miserocchi Stefano (ISMAR/CNR) - Zaccone Renata (IAMC/CNR) - Lavezza, R., Dubroca, L. F. C., Ludicone, D., Kress, N., Herut, B., Civitarese, G., Cruzado, A., Lefèvre, D.,Souvermezoglou, E., Yilmaz, A., Tugrul, S., and Ribera d'Alcala, M.: Compilation of quality controlled nutrient profiles from the Mediterranean Sea, doi:10.1594/PANGAEA.771907, 2011.
-

This visualization product displays the density of floating micro-litter per net normalized per km² per year from specific protocols different from research and monitoring protocols. EMODnet Chemistry included the collection of marine litter in its 3rd phase. Before 2021, there was no coordinated effort at the regional or European scale for micro-litter. Given this situation, EMODnet Chemistry proposed to adopt the data gathering and data management approach as generally applied for marine data, i.e., populating metadata and data in the CDI Data Discovery and Access service using dedicated SeaDataNet data transport formats. EMODnet Chemistry is currently the official EU collector of micro-litter data from Marine Strategy Framework Directive (MSFD) National Monitoring activities (descriptor 10). A series of specific standard vocabularies or standard terms related to micro-litter have been added to SeaDataNet NVS (NERC Vocabulary Server) Common Vocabularies to describe the micro-litter. European micro-litter data are collected by the National Oceanographic Data Centres (NODCs). Micro-litter map products are generated from NODCs data after a test of the aggregated collection including data and data format checks and data harmonization. A filter is applied to represent only micro-litter sampled according to a very specific protocol such as the Volvo Ocean Race (VOR) or Oceaneye. Densities were calculated for each net using the following calculation: Density (number of particles per km²) = Micro-litter count / (Sampling effort (km) * Net opening (cm) * 0.00001) When the number of microlitters or the net opening was not filled, it was not possible to calculate the density. Percentiles 50, 75, 95 & 99 have been calculated taking into account data for all years. Warning: the absence of data on the map does no't necessarily mean that they do not exist, but that no information has been entered in the National Oceanographic Data Centre (NODC) for this area.
-

This visualization product displays the density of floating micro-litter per net normalized in grams per km² per year from specific protocols different from research and monitoring protocols. EMODnet Chemistry included the collection of marine litter in its 3rd phase. Before 2021, there was no coordinated effort at the regional or European scale for micro-litter. Given this situation, EMODnet Chemistry proposed to adopt the data gathering and data management approach as generally applied for marine data, i.e., populating metadata and data in the CDI Data Discovery and Access service using dedicated SeaDataNet data transport formats. EMODnet Chemistry is currently the official EU collector of micro-litter data from Marine Strategy Framework Directive (MSFD) National Monitoring activities (descriptor 10). A series of specific standard vocabularies or standard terms related to micro-litter have been added to SeaDataNet NVS (NERC Vocabulary Server) Common Vocabularies to describe the micro-litter. European micro-litter data are collected by the National Oceanographic Data Centres (NODCs). Micro-litter map products are generated from NODCs data after a test of the aggregated collection including data and data format checks and data harmonization. A filter is applied to represent only micro-litter sampled according to a very specific protocol such as the Volvo Ocean Race (VOR) or Oceaneye. Densities were calculated for each net using the following calculation: Density (weight of particles per km²) = Micro-litter weight / (Sampling effort (km) * Net opening (cm) * 0.00001) When the weight of microlitters or the net opening was not filled, it was not possible to calculate the density. Percentiles 50, 75, 95 & 99 have been calculated taking into account data for all years. Warning: the absence of data on the map does not necessarily mean that they do not exist, but that no information has been entered in the National Oceanographic Data Centre (NODC) for this area.
-
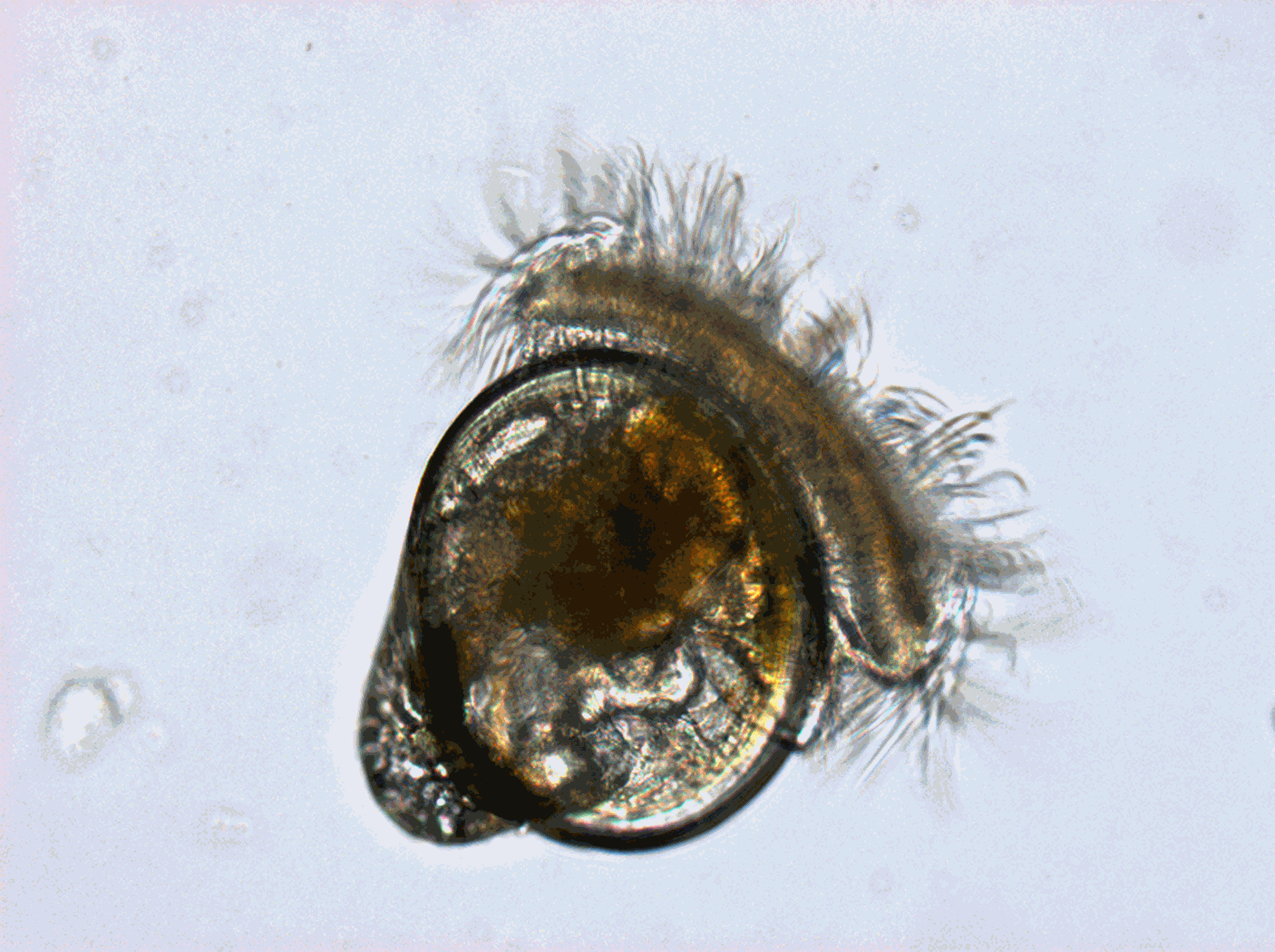
Worldwide, shellfish aquaculture and fisheries in coastal ecosystems represent crucial activities for human feeding. But these biological productions are under the pressure of climate variability and global change. Anticipating the biological processes affected by climate hazards remains a vital objective for species conservation strategies and human activities that rely on. Within marine species, filter feeders like oysters are real key species in coastal ecosystems due to their economic and societal value (fishing and aquaculture) but also due to their ecological importance. Indeed oysters populations in good health play the role of ecosystem engineers that can give many ecosystem services at several scales: building reef habitats that contribute to biodiversity, benthic-pelagic coupling and phytoplankton bloom control through water filtration, living shorelines against coastal erosion… The Pacific oyster, Crassostrea gigas (Thunberg, 1793), which is currently widespread worldwide, was introduced into the Atlantic European coasts at the end of the 19th century for shellfish culture purposes and becomes the main marine species farmed in France (around 100 000 tons) despite severe mortalities crisis. But in the same time and because of warming, natural oysters beds has spread significantly along the French coast and are supposed to have reach approximately 500 000 tons. In that context, Pacific oyster populations (natural and cultivated) in France are the subjects of many scientific projects. Among them, a specific long-term biological monitoring focuses on the reproduction of these populations at a national scale: the VELYGER national program. With more than 8 years of weekly data at many stations in France, this field-monitoring program offers a valuable dataset for studying processes underpinning reproduction cycle of this key-species in relation to environmental parameters, water quality and climate change. Database content: Larval concentration (number of individuals per 1.5 m3) monitored, since 2008, at several stations in six bays of the French coast (from south to north): Thau Lagoon and bays of Arcachon, Marennes Oléron, Bourgneuf, Vilaine and Brest (see map below). Methods used to monitor larval concentration: An important volume of seawater (1.5 m3) is pumped twice a week throughout the spawning season (june-september), at one meter below the surface at high tide (+/- 2h) in several sites within each VELYGER ecosystem. Water is filtered trough plankton net fitted with 40 µm mesh. After a proper rinsing of the net, the retained material is transferred into a polyethylene bottle (1 liter) and fixed with alcohol. At laboratory, sample is then gently filtered and rinse again and transferred into eprouvette. Two sub-samples of 1 mL are then taken using a pipette and examined on a graticule slide for microscope. The microscopic examination is made with a conventional binocular optical microscope with micrometer stage at a magnification of 10 X (or above). During the counting, a special care is necessary as larvae of other bivalves are also collected and confusion is possible. Larvae of C. gigas are also classified into four stage of development: - Stage I = D-shaped straight hinge larvae (shell length <105 µm) - Stage II = Early umbo evolved larvae (shell length between 105 and 150 µm) - Stage III = Medium umbo larvae (shell length between 150 and 235 µm) - Stage IV*= Large umbo eyed pediveliger larvae (shell length > 235 µm) * Larvae that are very closed to settle are sometimes identified into a separated 5th stage, but generally this stage is included in stage IV. Illustrations: Location of the different Velyger sites along the French coast. From south to north: Thau Lagoon and bays of Arcachon, Marennes Oléron, Bourgneuf, Vilaine and Brest. Legend: Pacific Oyster Larvae (left side) and Natural oyster bed (right side). Photos : © S. Pouvreau/Ifremer
-

Particularly suited to the purpose of measuring the sensitivity of benthic communities to trawling, a trawl disturbance indicator (de Juan and Demestre, 2012, de Juan et al. 2009) was proposed based on benthic species life history traits to evaluate the sensibility of mega- and epifaunal community to fishing pressure known to have a physical impact on the seafloor (such as dredging and bottom trawling). The selected biological traits were chosen as they determine vulnerability to trawling: mobility, fragility, position on substrata, average size and feeding mode that can easily be related to the fragility, recoverability and vulnerability ecological concepts. Life history traits of species have been defined from the BIOTIC database (MARLIN, 2014) and from information given by Le Pape et al. (2007), Brindamour et al. (2009) and Garcia (2010). For missing life history traits, additional information from literature has been considered. The five categories retained are life history functional traits that were selected based on the knowledge of the response of benthic taxa to trawling disturbance (de Juan and Demestre, 2012). They reflect respectively the possibility to avoid direct gear impact, to benefit from trawling for feeding, to escape gear, to get caught by the net and to resist trawling/dredging action, each of these characteristics being either advantageous or sensitive to trawling. Then, to allow quantitative analysis, a score was assigned to each category: from low vulnerability (0) to high vulnerability (3). The five categories scores were then summed for each taxon (the highly vulnerable taxon could reach the maximum score is 15) and this value may be considered as a species index of sensitivity to trawling disturbance. The scores of 812 taxa commonly found in bottom trawl by-catch in the southern North Sea, English Channel and north-western Mediterranean were described.
-

The ARCHYD dataset, which have been collected since 1988, represents the longest long-term hydrologic data sets in Arcachon Bay. The objectives of this monitoring programme are to assess the influence of oceanic and continental inputs on the water quality of the bay and their implications on biological processes. It also aims to estimate the effectiveness of management policies in the bay by providing information on trends and/or shifts in pressure, state, and impact variables. Sampling is carried on stations spread across the entire bay, but since 1988, the number and location of stations have changed slightly to better take into account the gradient of ocean and continental inputs. In 2005, the ARCHYD network was reduced to 8 stations that are still sampled by Ifremer to date. All the stations are sampled at a weekly frequency, at midday, alternately around the low spring tide and the high neap tide. Data are complementary to REPHY dataset. Physico-chemical measures include temperature, salinity, turbidity, suspended matters (organic, mineral), dissolved oxygen and dissolved inorganic nutrients (ammonium, nitrite+nitrate, phosphate, silicate). Biological measures include pigment proxies of phytoplankton biomass and state (chlorophyll a and phaeopigment).
 Catalogue PIGMA
Catalogue PIGMA